Research Grants & Tools
Our research is supported by the NSF CAREER award (DBI-1652164), the NIH R01 award (R01GM140370), and a University of Nebraska–Lincoln startup grant.


Bioinformatics Tool Development
ORFan study (NIH 1R15GM114706): We develop bioinformatics tools for studying orphan genes and horizontally transferred genes in plant, microbial, and metagenomic datasets.
• ORFanFinder identifies taxonomically restricted genes and classifies genes into evolutionary age groups. Visit ORFanFinder
• HGT-Finder identifies horizontally transferred genes in fungi.

CAZyme Annotation & Databases
CAZyme study (NSF DBI-1652164, NIH R01GM140370): We develop automated pipelines for genome-scale CAZyme annotation.
• dbCAN — a domain-based CAZyme annotation resource.
• PlantCAZyme — precomputed CAZyme annotations for all sequenced plant and algal genomes.


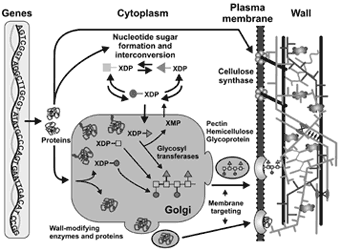
Evolution of Plant Cell Walls
We sequence and analyze charophyte green algae such as Zygnema circumcarinatum to understand the evolutionary transition from algae to land plants.
We study phylogenies of cell wall-related genes encoding enzymes, transcription factors, transporters, and ncRNAs involved in synthesis and degradation.
We identify industrially important enzymes by large-scale mining of genomic and metagenomic datasets.