📊 MAG Datasets of Human Gut Microbiomes
- This table summarizes the metagenome-assembled genomes (MAGs) from diverse human populations and lifestyles that are used across the platform.
| Database | Reference | Populations/Lifestyles | # of MAGs Used | # in Non-Redundant Set | # Representative |
|---|---|---|---|---|---|
| H3Africa Phase 1 | Tamburini et al., 2022 | 190 women in South Africa | 817 | 807 | 3 |
| H3Africa Phase 2 | Maghini et al., 2025 | 1,820 adults across 4 African countries | 1796 | 1779 | 706 |
| Hadza | Carter et al., 2024 | 167 Hadza hunter-gatherers | 961 | 960 | 682 |
| UHGG | Almeida et al., 2021 | Mostly Global North countries | 2424 | 1980 | 1055 |
| WIS | Leviatan et al., 2022 | Mostly Israel and USA | 2173 | 1540 | 608 |
| ELGG | Zeng et al., 2022 | Children under 3 (Global North) | 315 | 299 | 178 |
| CGMR | Huang et al., 2024 | Chinese Gut Microbial Reference | 2024 | 1989 | 811 |
| HRGM2 | Ma et al., 2024 | 41 countries (Asia, Africa, S. America) | 2324 | 2307 | 1743 |
| IMGG | Jin et al., 2023 | Inner Mongolia (ONT + Illumina) | 481 | 478 | 206 |
| SPMP | Gounot et al., 2022 | 109 healthy Singaporeans | 1769 | 1747 | 39 |
| Total | 15,084 | 13,886 | 6,031 |
🏠 Home Page
- Quickly browse and search genomes, CGCs, and CGC families with ease.
🧬 Genome Page
- Metadata and basic information of genomes can be downloaded after genome browsing and searching.
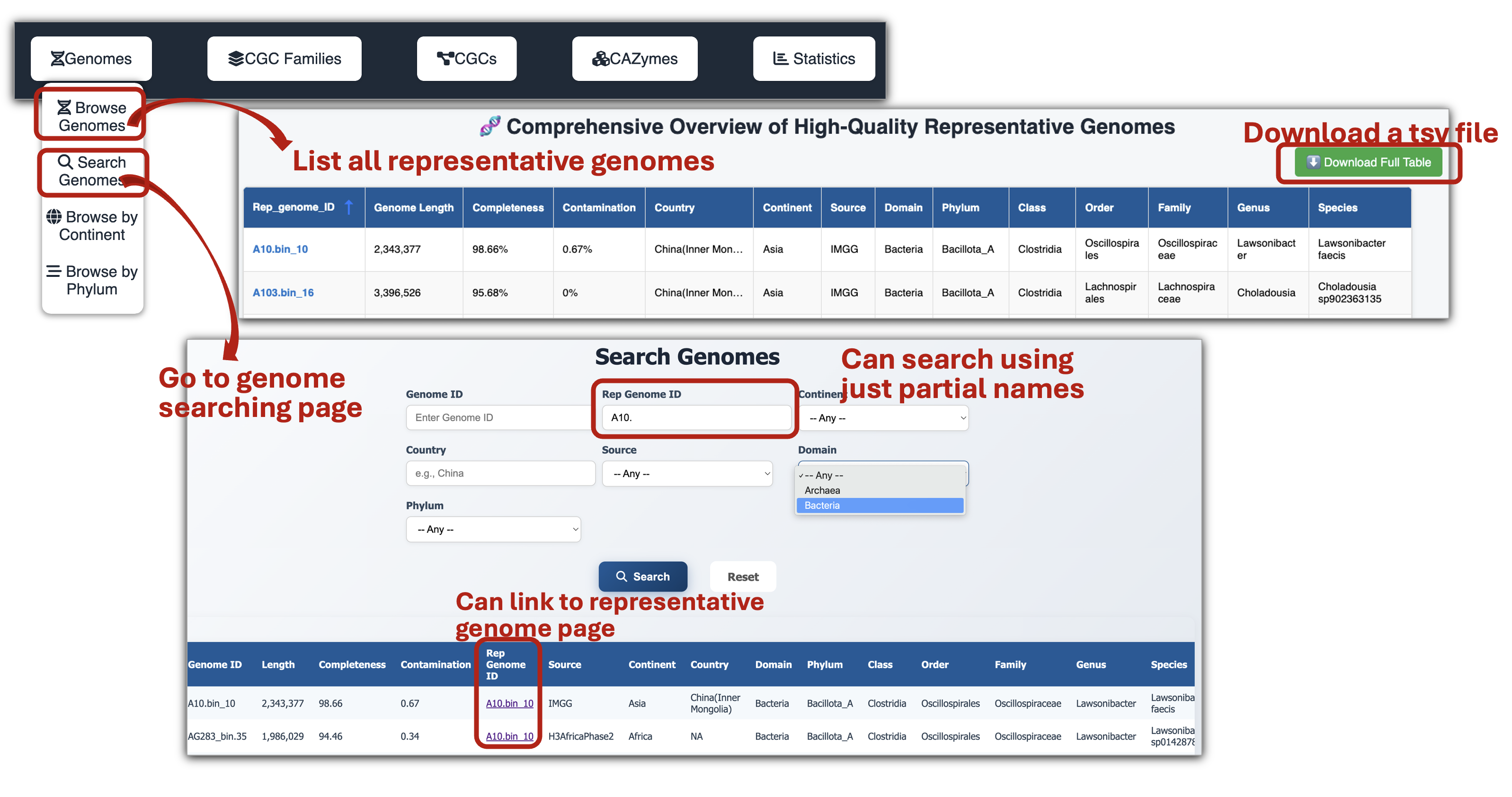
- Each representative genome page provides the Genome list, Basic statistics plots, Taxonomy lineage, CGC list of the rep genome, as well as Read mapping on CGC regions visualized by JBrowse.

🧪 CGC Family Page
- Browse CGC families by Substrate to view a list of families and explore the top abundant CAZyme families within each substrate through interactive plots.
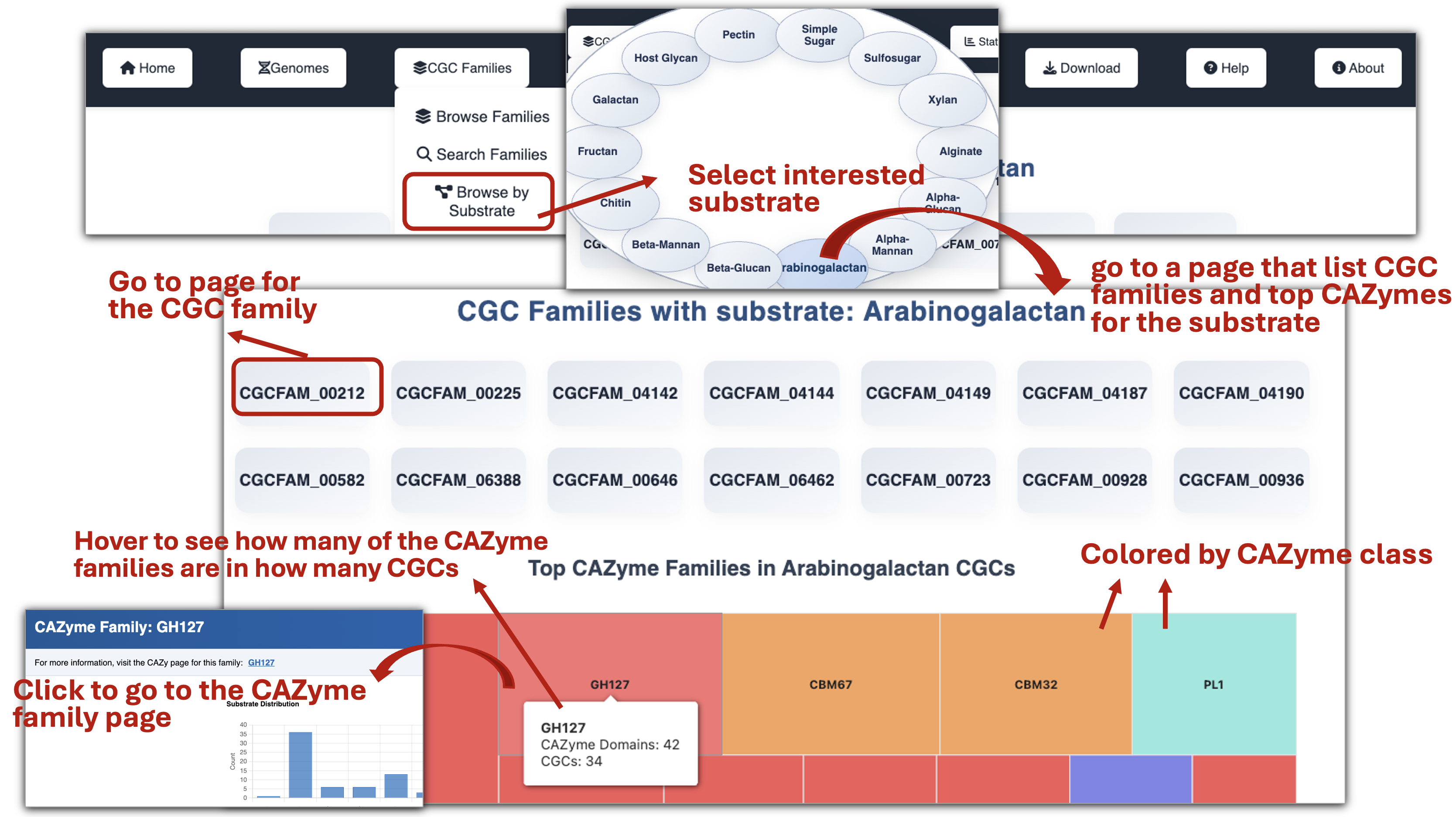
- Browse all CGC families to explore the substrate information of their CGCs and access links to each family’s detailed page.
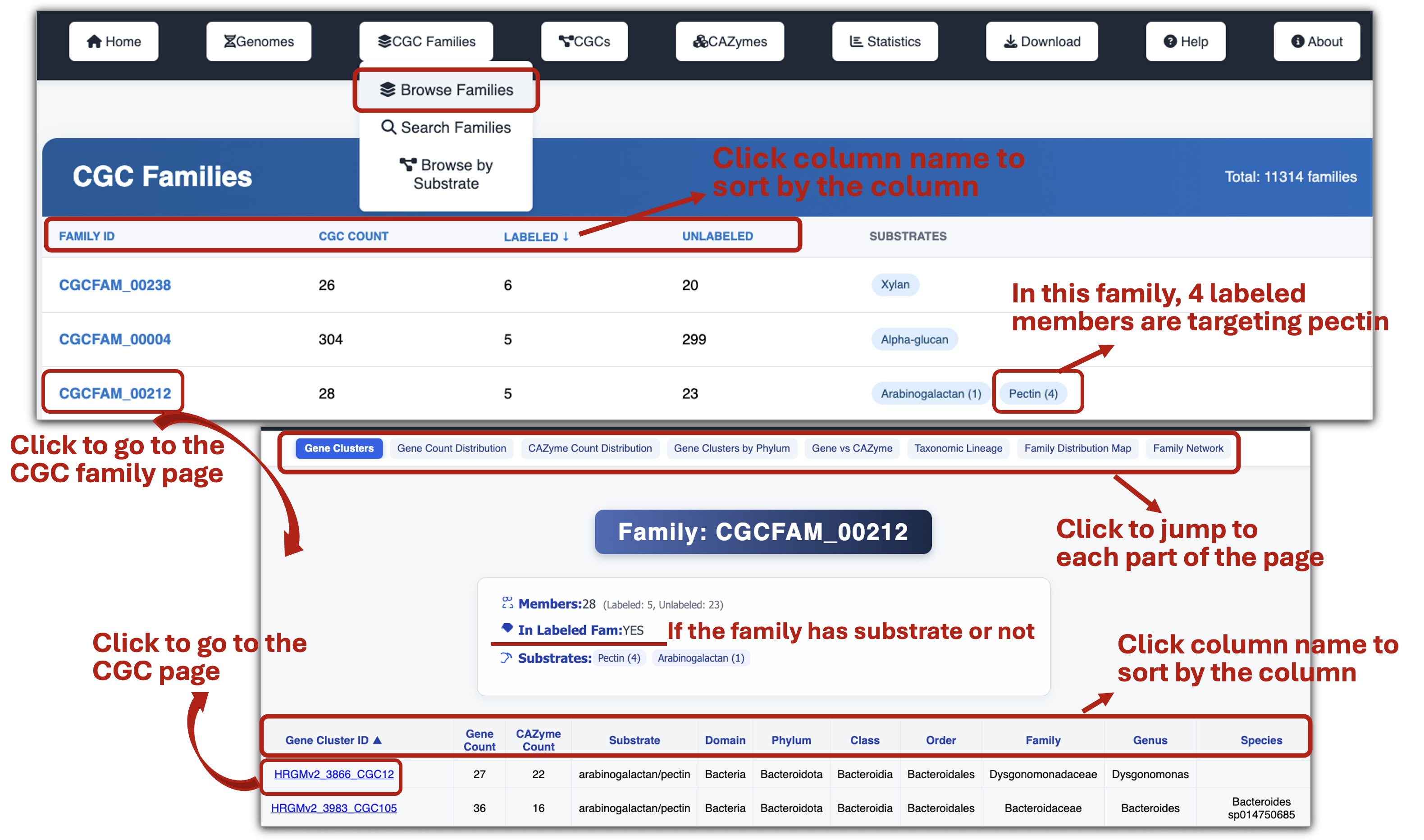
- Each CGC family page provides basic statistics plots, taxonomy distribution plot and network diagram of the family members.
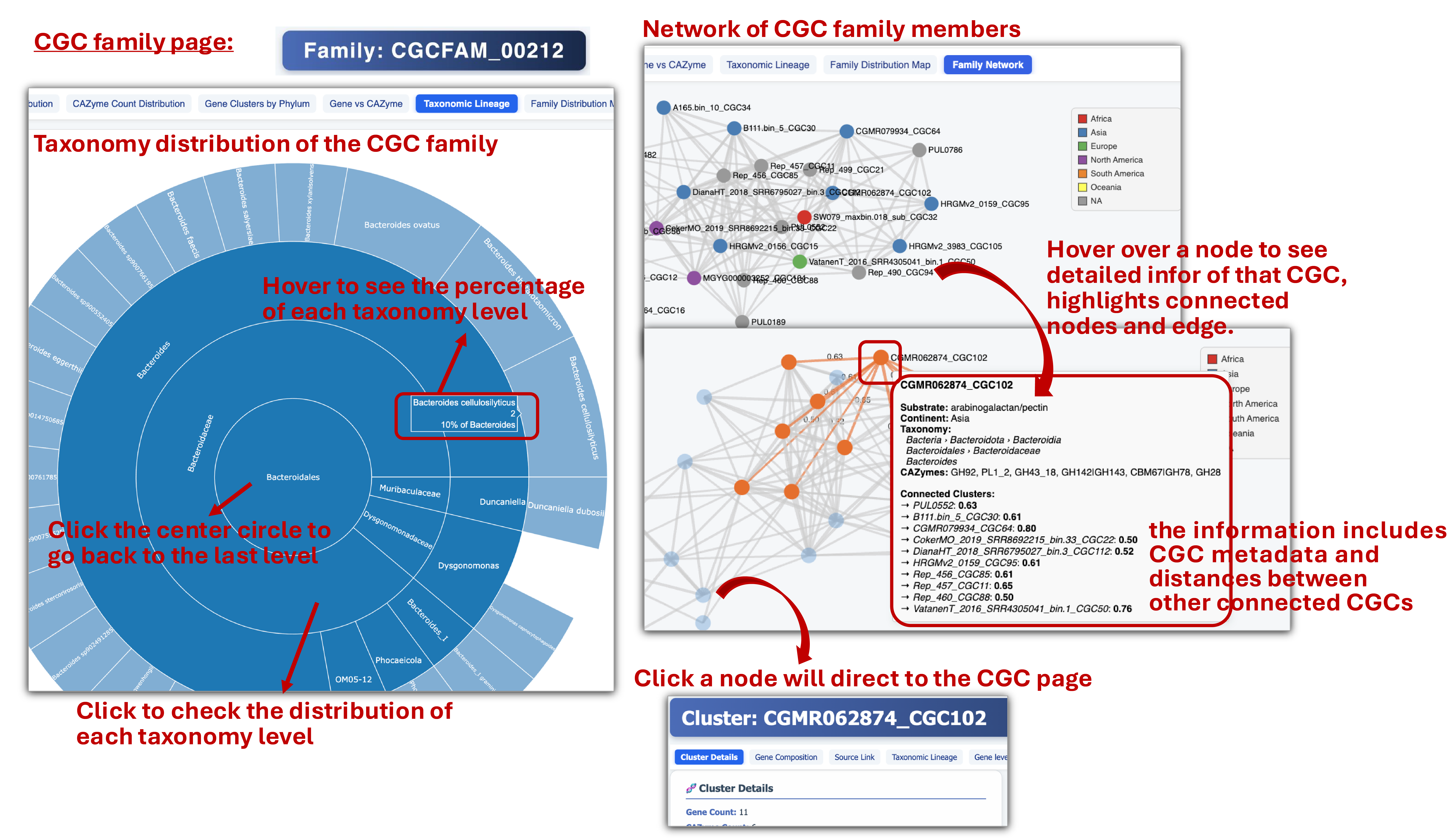
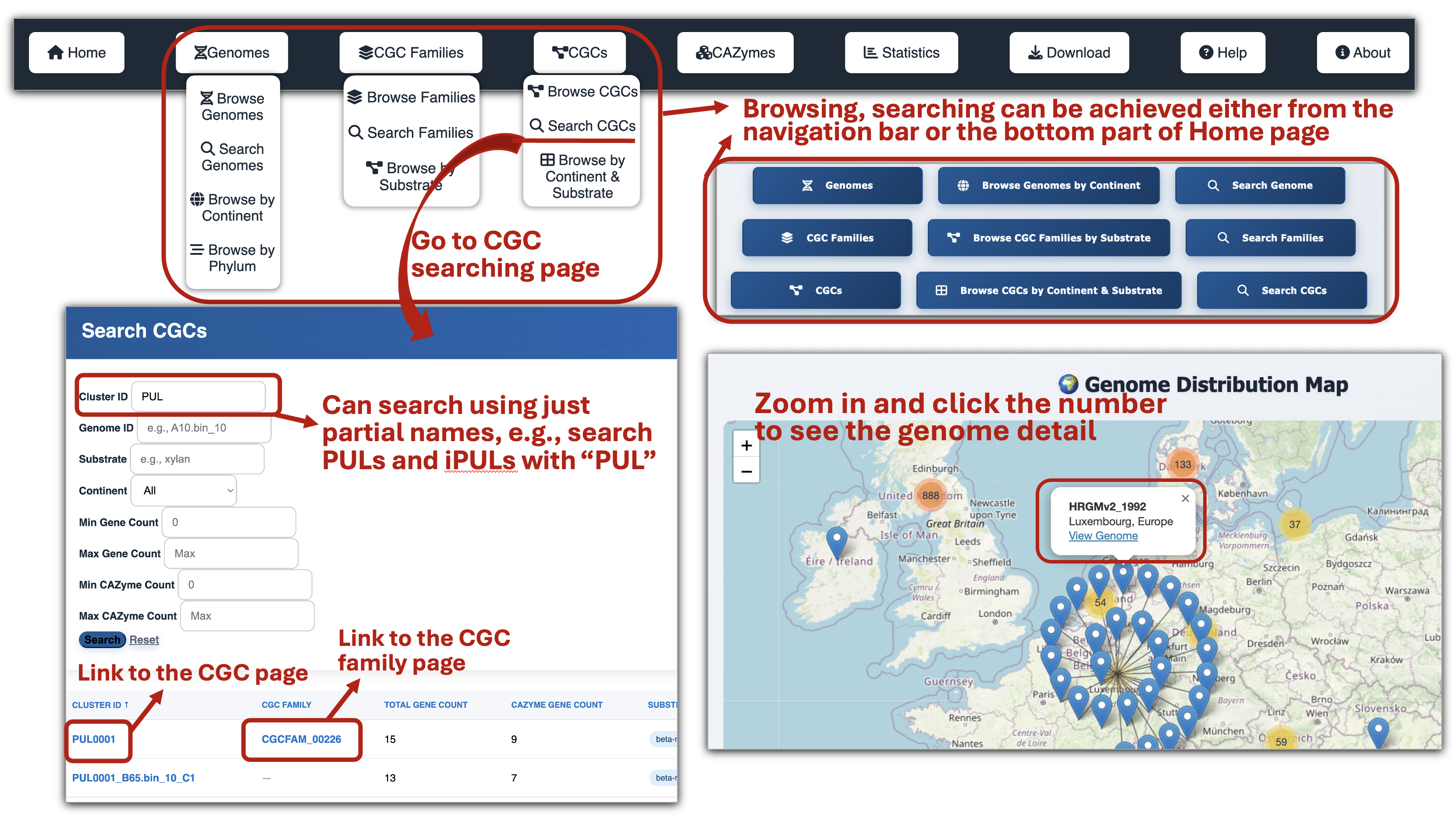
🧬 CGC (Gene Cluster) Page
- Browse CGCs by continent and substrate to explore the distribution of CGCs targeting various substrates across different continents.
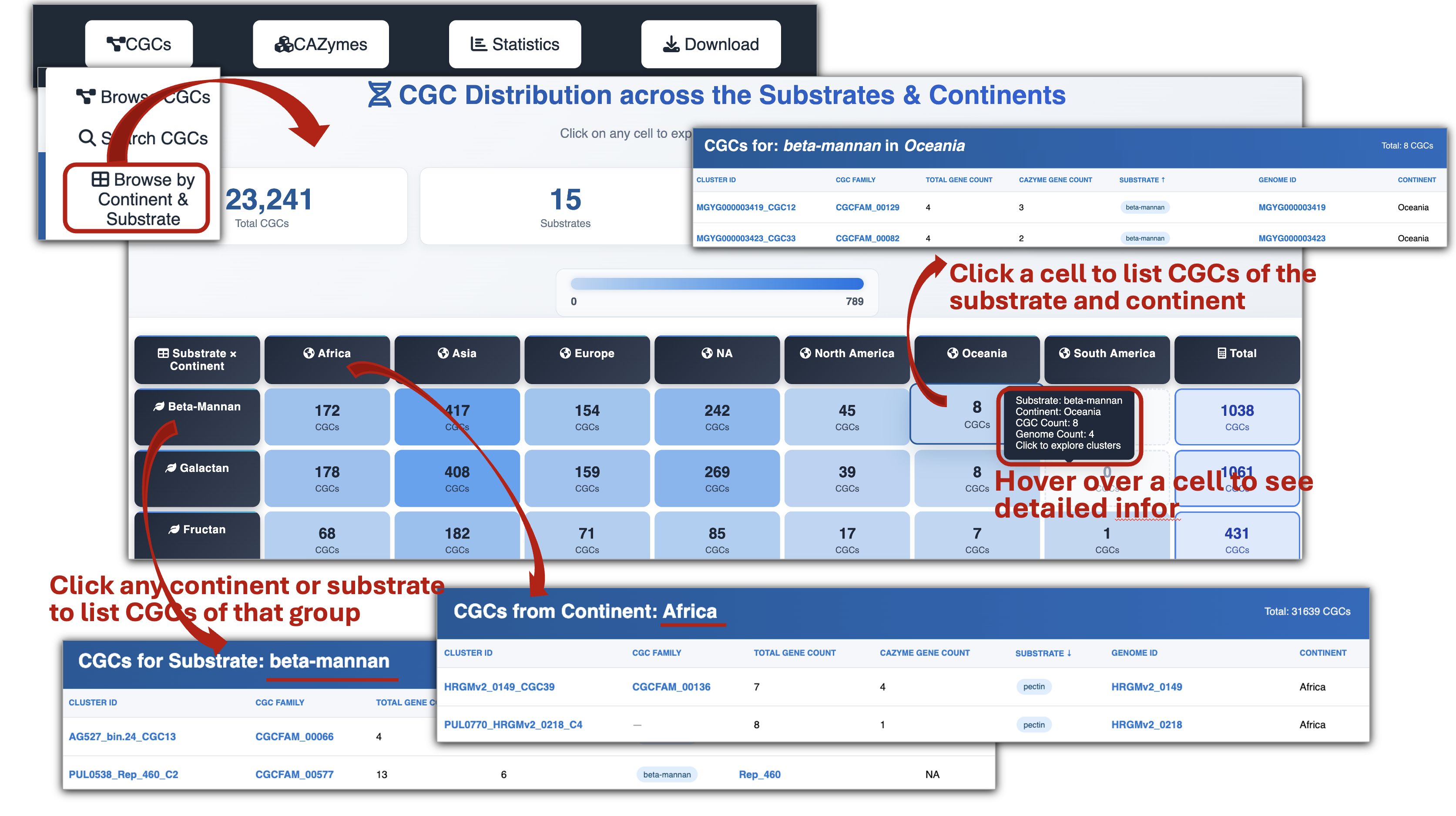
- Each CGC page provides a Gene composition table, links to Protein gene annotation resources, and information on Genome source and Taxonomic lineage.
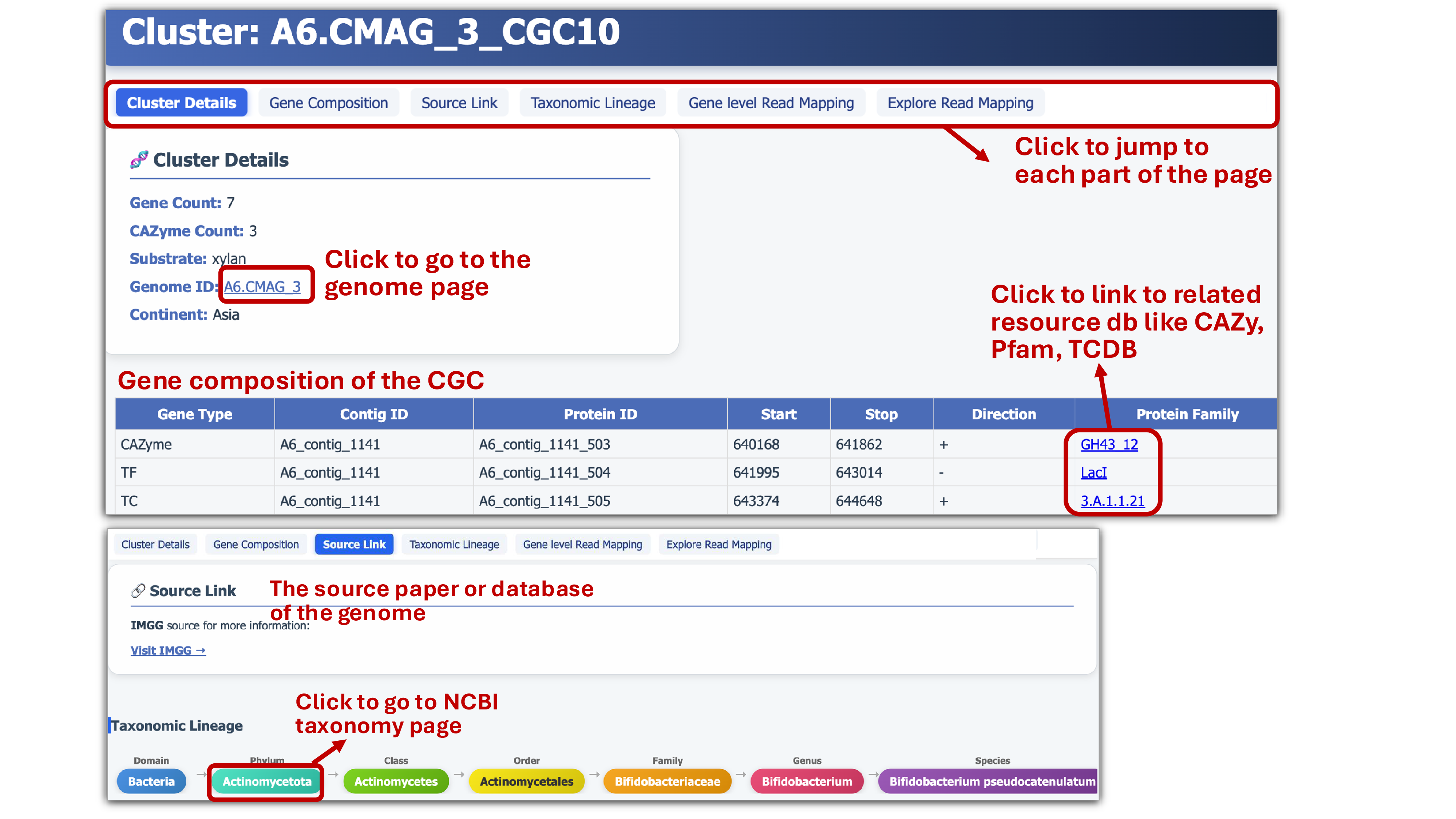
- The null genes without sequence homology annotation are linked to a protein structure page, containing predicted 3D structure visualization with pLDDT confidence scores and results of structural homologs against AFDB, CAZyme3D, CAZymeID50, PDB, and SwissProt.

- Each CGC page also includes Read coverage plots from diet intervention studies to compare CGC coverage in gut metagenome samples across different diets, along with a gene composition diagram.
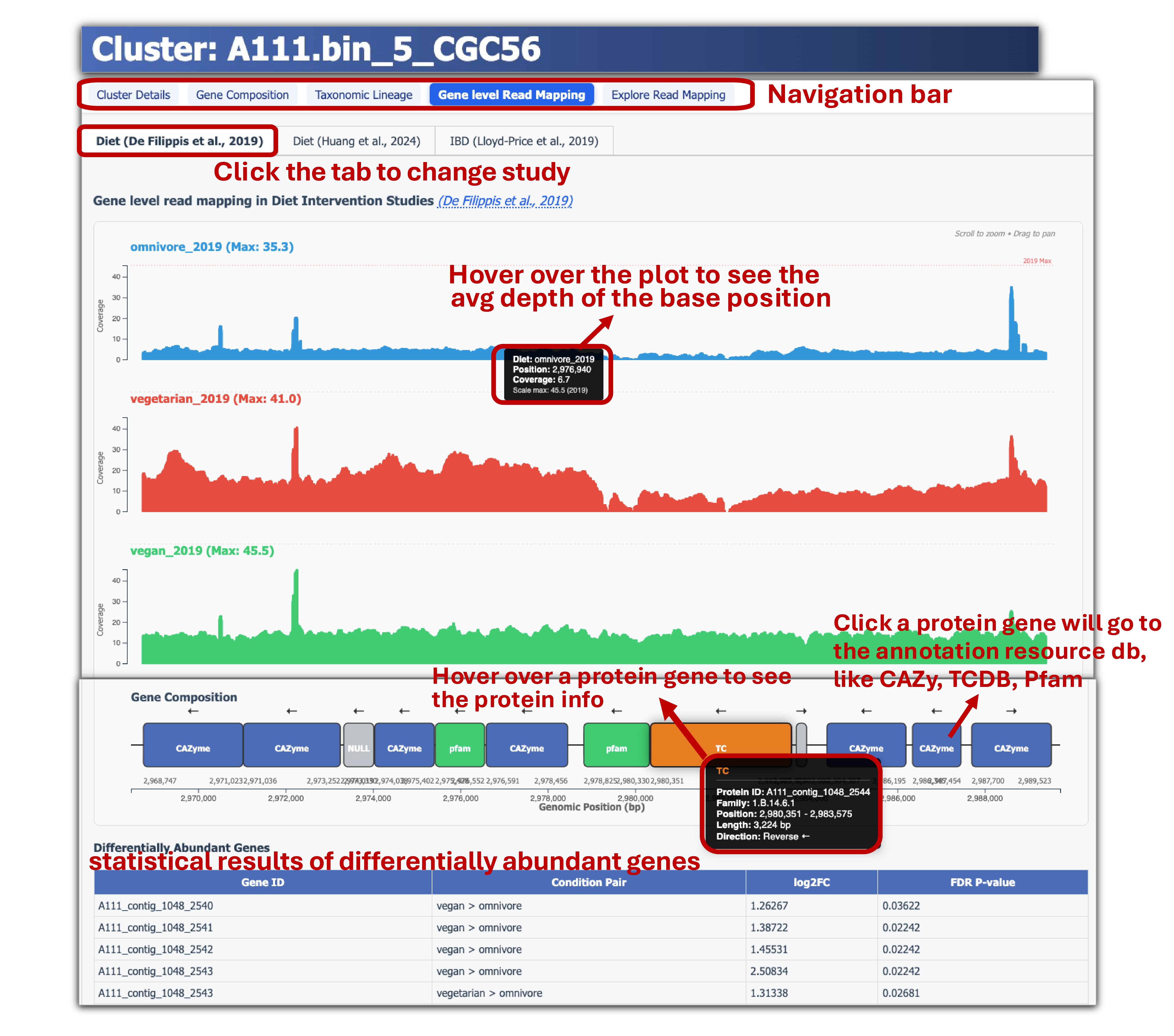
- Explore read mapping of CGC across individual samples using JBrowse.

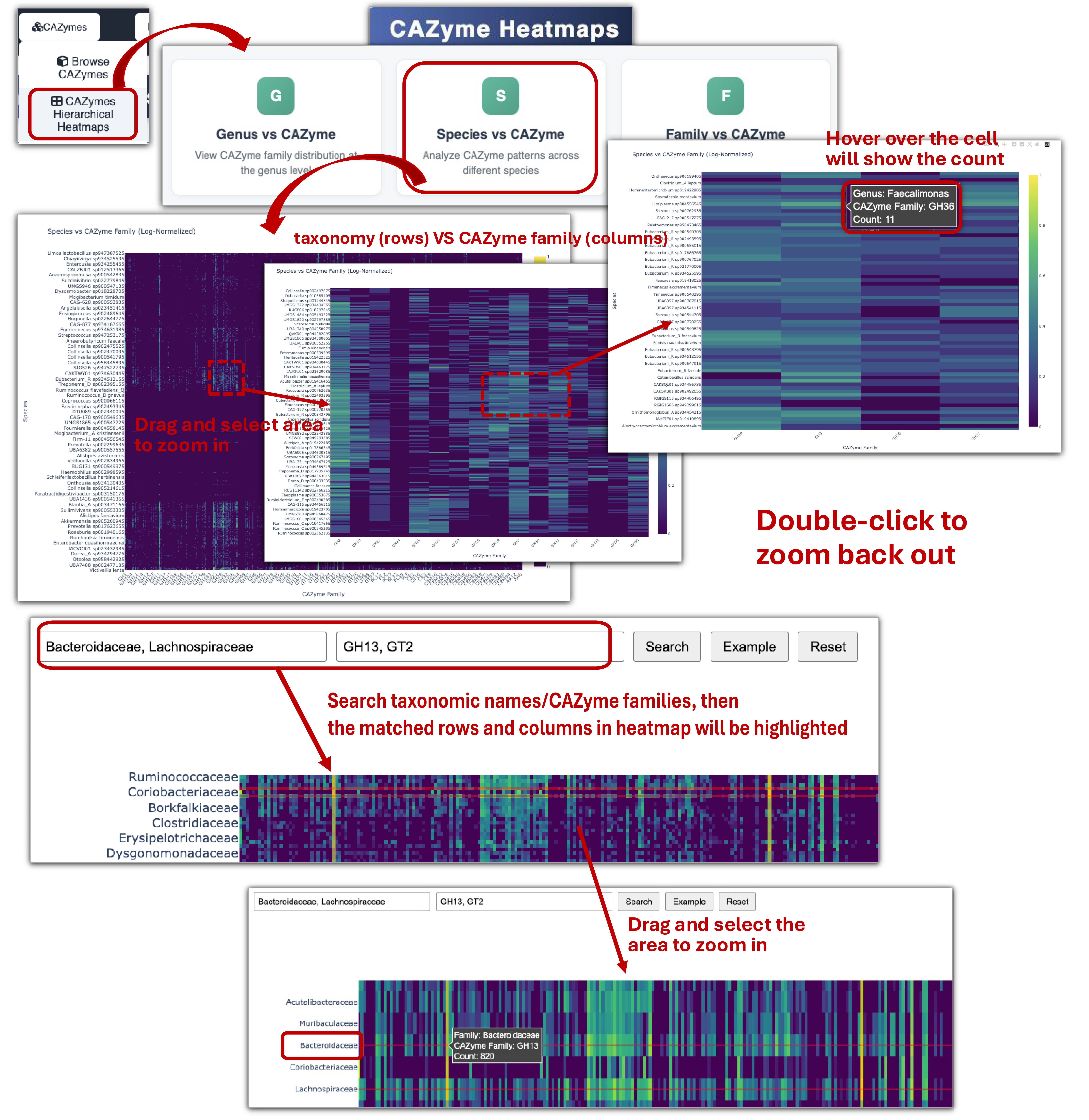
🗺️ CAZymes HeatMap
- Interactive heatmap displaying the distribution of CAZyme families in CGCs across species and genera. Users can select regions of the heatmap to zoom in for a more detailed view and exploration of specific family abundances.