Seed proteins |
Seed genomic context | Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS067 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | ||||||||||||||
| CLAN ID | CLAN003 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | ||||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | PF07275.14 | |||||||||||||
| Phrog | phrog_2059 | |||||||||||||
| Host taxa | ||||||||||||||
| Gene Location | Start: 6069; End: 6716; Strand: + | |||||||||||||
| Description | ||||||||||||||
| Note | This family does not have seed protein, but we infer its function is defending restriction-modification (RM) because it belongs to the same clan as family APIS003,ardu which has seed protein. | |||||||||||||
Seed genomic context Download table help
| Genome accession | Start | End | Strand | Protein ID | Length | Molecular weight | Charge | Isoelectric point | Function | Phrog | PFAM |
|---|---|---|---|---|---|---|---|---|---|---|---|
| IMGVR_UViG_3300035506_000204 | 10713 | 11669 | + | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_10713_11669 | 318 | 35543.45 | -8.0 | 4.7897 | hypothetical protein | ||
| IMGVR_UViG_3300035506_000204 | 11770 | 12363 | + | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_11770_12363 | 197 | 21122.49 | -6.5 | 4.4764 | hypothetical protein | ||
| IMGVR_UViG_3300035506_000204 | 3654 | 3965 | + | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_3654_3965 | 103 | 11771.41 | -1.0 | 5.2600 | DNA-binding XRE family transcriptional regulator | phrog_38027 ; phrog_1373 ; phrog_2439 ; phrog_3 ; phrog_29047 ; phrog_6831 ; phrog_11553 ; phrog_24325 ; phrog_5823 ; phrog_2339 ; phrog_1381 | PF01381.25 |
| IMGVR_UViG_3300035506_000204 | 4146 | 4622 | + | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_4146_4622 | 158 | 19114.97 | 6.0 | 8.9352 | hypothetical protein | ||
| IMGVR_UViG_3300035506_000204 | 4632 | 4757 | + | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_4632_4757 | 41 | 4601.38 | -5.5 | 3.8121 | hypothetical protein | ||
| IMGVR_UViG_3300035506_000204 | 4760 | 5188 | + | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_4760_5188 | 142 | 15933.44 | 1.5 | 7.1875 | hypothetical protein | ||
| IMGVR_UViG_3300035506_000204 | 5461 | 5832 | - | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_5461_5832 | 123 | 14044.42 | -2.0 | 5.1657 | hypothetical protein | ||
| IMGVR_UViG_3300035506_000204 | 6069 | 6716 | + | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_6069_6716 | 215 | 25113.60 | -29.0 | 3.9397 | antirestriction protein | phrog_2059 | PF07275.14 |
| IMGVR_UViG_3300035506_000204 | 6790 | 7458 | - | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_6790_7458 | 222 | 26369.84 | 0.5 | 6.6193 | hypothetical protein | ||
| IMGVR_UViG_3300035506_000204 | 7698 | 9344 | - | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_7698_9344 | 548 | 62551.21 | -6.0 | 5.1316 | hypothetical protein | phrog_8741 | PF08238.15 |
| IMGVR_UViG_3300035506_000204 | 7698 | 9344 | - | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_7698_9344 | 548 | 62551.21 | -6.0 | 5.1316 | hypothetical protein | phrog_8741 | PF08238.15 |
| IMGVR_UViG_3300035506_000204 | 7698 | 9344 | - | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_7698_9344 | 548 | 62551.21 | -6.0 | 5.1316 | hypothetical protein | phrog_8741 | PF08238.15 |
| IMGVR_UViG_3300035506_000204 | 7698 | 9344 | - | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_7698_9344 | 548 | 62551.21 | -6.0 | 5.1316 | hypothetical protein | phrog_8741 | PF08238.15 |
| IMGVR_UViG_3300035506_000204 | 7698 | 9344 | - | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_7698_9344 | 548 | 62551.21 | -6.0 | 5.1316 | hypothetical protein | phrog_8741 | PF08238.15 |
| IMGVR_UViG_3300035506_000204 | 7698 | 9344 | - | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_7698_9344 | 548 | 62551.21 | -6.0 | 5.1316 | hypothetical protein | phrog_8741 | PF08238.15 |
| IMGVR_UViG_3300035506_000204 | 9450 | 10571 | + | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_9450_10571 | 373 | 40537.68 | -11.5 | 4.7223 | ATP-dependent protease ClpP protease subunit | phrog_94 ; phrog_1684 ; phrog_3348 ; phrog_133 ; phrog_2976 ; phrog_53 | PF00574.26 |
Seed genomic context- JBrowse
Predicted 3D structure by alphafold2 with pTM = 0.82 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
Structure homologs against by afdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| AF-A0A0H3GZQ3-F1-model_v4 | 49 | 214 | 5 | 168 | 0.8453 | 0.7359 |
Structure homologs against by esmdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| MGYP002619016300.pdb | 6 | 215 | 1 | 211 | 0.9465 | 0.9277 |
| MGYP003538369595.pdb | 28 | 215 | 1 | 186 | 0.8888 | 0.8145 |
| MGYP000693494289.pdb | 4 | 215 | 2 | 205 | 0.8558 | 0.8121 |
| MGYP002521666680.pdb | 8 | 215 | 1 | 200 | 0.8541 | 0.8009 |
| MGYP002519753648.pdb | 21 | 215 | 1 | 188 | 0.8717 | 0.7998 |
| MGYP002523803474.pdb | 21 | 215 | 16 | 207 | 0.8568 | 0.7947 |
| MGYP003228289826.pdb | 28 | 215 | 2 | 190 | 0.8598 | 0.7937 |
| MGYP003324507119.pdb | 18 | 215 | 14 | 205 | 0.8442 | 0.7931 |
| MGYP003510632526.pdb | 24 | 215 | 1 | 190 | 0.8578 | 0.79 |
| MGYP003439714399.pdb | 12 | 215 | 1 | 208 | 0.8281 | 0.7883 |
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS067 | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_6069_6716 | 18 | HMM model | Member alignment |
|---|
Host Taxa distribution
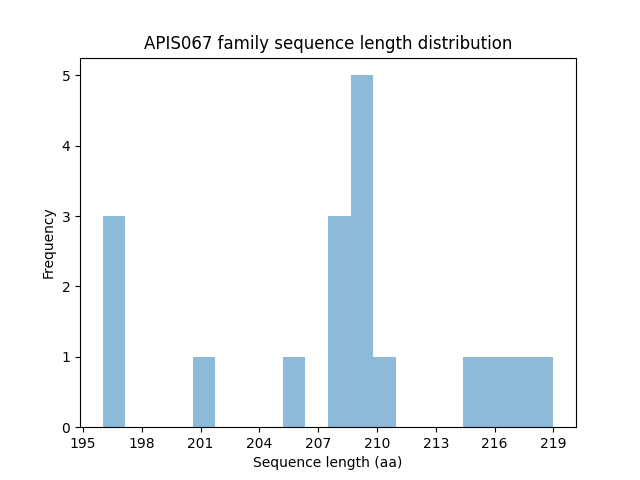
Length distribution
Homology to other families download full data without filtering help
Clan ID: CLAN003
| Hit APIS family | E-Value | Query Start | Query End | Hit Start | Hit End | Immune system | Seed protein |
|---|---|---|---|---|---|---|---|
| APIS003 | 4.57E-69 | 48 | 215 | 36 | 203 | restriction-modification (RM) | AAB36891.1 |
| APIS063 | 5.95E-54 | 46 | 215 | 8 | 174 | - | IMGVR_UViG_3300031923_000006|3300031923|Ga0326331_1000015821 |
| APIS051 | 9.64E-54 | 48 | 214 | 54 | 222 | - | IMGVR_UViG_3300012016_000701|3300012016|Ga0120387_100082722 |
| APIS072 | 2.75E-41 | 47 | 214 | 48 | 223 | - | IMGVR_UViG_3300039331_000150|3300039331|Ga0169804_01911_10006_10671 |
