Seed proteins |
Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS051 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | ||||||||||||||
| CLAN ID | CLAN003 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | ||||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | PF07275.14 | |||||||||||||
| Phrog | phrog_14321,phrog_2059 | |||||||||||||
| Host taxa | ||||||||||||||
| Gene Location | Start: ; End: ; Strand: | |||||||||||||
| Description | ||||||||||||||
| Note | This family does not have seed protein, but we infer its function is defending restriction-modification (RM) because it belongs to the same clan as family APIS003,ardu which has seed protein. | |||||||||||||
Predicted 3D structure by alphafold2 with pTM = 0.7 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
Structure homologs against by afdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| AF-A0A0H3GZQ3-F1-model_v4 | 55 | 223 | 5 | 168 | 0.7952 | 0.6754 |
Structure homologs against by esmdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| MGYP003228289826.pdb | 37 | 223 | 1 | 190 | 0.9258 | 0.8467 |
| MGYP002627441165.pdb | 36 | 221 | 1 | 189 | 0.9162 | 0.8361 |
| MGYP003538369595.pdb | 39 | 223 | 1 | 186 | 0.9113 | 0.8242 |
| MGYP002522264593.pdb | 29 | 222 | 2 | 195 | 0.8904 | 0.8233 |
| MGYP002523803474.pdb | 41 | 223 | 23 | 207 | 0.9271 | 0.8031 |
| MGYP002855897325.pdb | 34 | 223 | 34 | 223 | 0.944 | 0.7996 |
| MGYP003306129017.pdb | 27 | 222 | 10 | 202 | 0.8456 | 0.7918 |
| MGYP002624915303.pdb | 20 | 223 | 9 | 205 | 0.8283 | 0.7829 |
| MGYP000232002069.pdb | 55 | 222 | 2 | 166 | 0.9078 | 0.7805 |
| MGYP003298723900.pdb | 54 | 222 | 1 | 170 | 0.9003 | 0.78 |
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS051 | IMGVR_UViG_3300012016_000701|3300012016|Ga0120387_100082722 | 8 | HMM model | Member alignment |
|---|
Host Taxa distribution
Length distribution
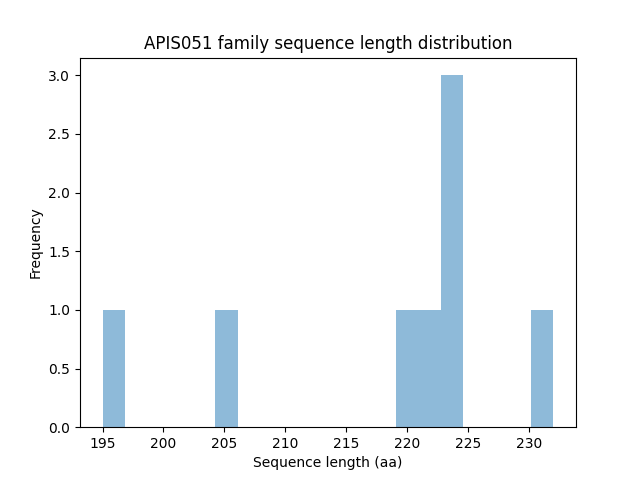
Homology to other families download full data without filtering help
Clan ID: CLAN003
| Hit APIS family | E-Value | Query Start | Query End | Hit Start | Hit End | Immune system | Seed protein |
|---|---|---|---|---|---|---|---|
| APIS003 | 1.95E-67 | 54 | 222 | 36 | 202 | restriction-modification (RM) | AAB36891.1 |
| APIS067 | 5.57E-55 | 45 | 222 | 32 | 206 | - | IMGVR_UViG_3300035506_000204|3300035506|Ga0376461_0000200_6069_6716 |
| APIS063 | 4.49E-54 | 50 | 222 | 6 | 173 | - | IMGVR_UViG_3300031923_000006|3300031923|Ga0326331_1000015821 |
| APIS072 | 2.37E-41 | 55 | 222 | 50 | 223 | - | IMGVR_UViG_3300039331_000150|3300039331|Ga0169804_01911_10006_10671 |
