Seed proteins |
Seed genomic context | Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS035 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | broad-spectrum counter-defense | |||||||||||||
| CLAN ID | CLAN032 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | Silas et al., 2023 | |||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | ||||||||||||||
| Phrog | phrog_3972,phrog_16544 | |||||||||||||
| Host taxa | d__Bacteria;p__Proteobacteria;c__Gammaproteobacteria;o__Enterobacterales;f__Enterobacteriaceae;g__Escherichia; | |||||||||||||
| Gene Location | Start: 6432; End: 6779; Strand: - | |||||||||||||
| Description | Broad-spectrum counter-defense. | |||||||||||||
Seed genomic context Download table help
| Genome accession | Start | End | Strand | Protein ID | Length | Molecular weight | Charge | Isoelectric point | Function | Phrog | PFAM |
|---|---|---|---|---|---|---|---|---|---|---|---|
| NC_047835.1 | 4114 | 4467 | - | YP_009790933.1 | 117 | 13141.82 | -8.0 | 4.2178 | hypothetical protein | phrog_7572 | |
| NC_047835.1 | 4557 | 4700 | - | YP_009790934.1 | 47 | 5481.57 | -1.0 | 4.5440 | hypothetical protein | phrog_13931 | |
| NC_047835.1 | 4700 | 5017 | - | YP_009790935.1 | 105 | 12702.88 | 5.0 | 8.0941 | hypothetical protein | phrog_6053 | |
| NC_047835.1 | 5115 | 5282 | - | YP_009790936.1 | 55 | 6458.46 | 5.5 | 10.0624 | hypothetical protein | phrog_6221 | |
| NC_047835.1 | 5469 | 5627 | - | YP_009790937.1 | 52 | 5856.22 | 2.5 | 8.5537 | hypothetical protein | phrog_18174 | |
| NC_047835.1 | 6432 | 6779 | - | YP_009790938.1 | 115 | 13375.95 | -12.0 | 4.0052 | hypothetical protein | phrog_16544 ; phrog_3972 | |
| NC_047835.1 | 6892 | 7077 | - | YP_009790939.1 | 61 | 7159.32 | 1.0 | 7.2032 | hypothetical protein | phrog_5473 | |
| NC_047835.1 | 7332 | 7523 | - | YP_009790940.1 | 63 | 7249.18 | -1.0 | 5.6351 | hypothetical protein | phrog_7042 | |
| NC_047835.1 | 7524 | 7742 | - | YP_009790941.1 | 72 | 7920.04 | -4.0 | 4.2850 | hypothetical protein | phrog_8408 | |
| NC_047835.1 | 7793 | 8065 | - | YP_009790942.1 | 90 | 10828.58 | 11.0 | 10.1441 | hypothetical protein | phrog_5876 | |
| NC_047835.1 | 8543 | 8818 | - | YP_009790943.1 | 91 | 10200.65 | -10.0 | 3.9759 | hypothetical protein | phrog_4991 ; phrog_4551 |
Seed genomic context- JBrowse
Predicted 3D structure by alphafold2 with pTM = 0.8 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
No homologs found in AlphaFold database
Structure homologs against by esmdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| MGYP000932118009.pdb | 2 | 115 | 1 | 98 | 0.6662 | 0.63 |
| MGYP003611076310.pdb | 1 | 115 | 1 | 96 | 0.6362 | 0.5643 |
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS035 | YP_009790938.1 | 111 | HMM model | Member alignment |
|---|
Host Taxa distribution
Length distribution
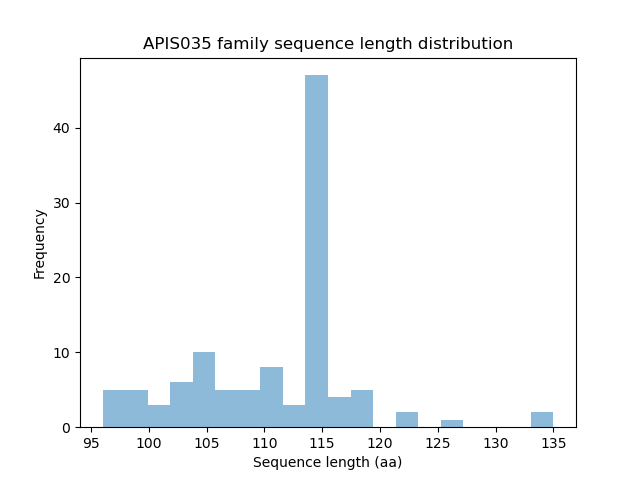
Homology to other families download full data without filtering help
Clan ID: CLAN032
| Hit APIS family | E-Value | Query Start | Query End | Hit Start | Hit End | Immune system | Seed protein |
|---|---|---|---|---|---|---|---|
| APIS075 | 6.32E-55 | 1 | 97 | 26 | 129 | - | KY083726_00051 |
| APIS047 | 7.98E-54 | 1 | 98 | 21 | 124 | - | IMGVR_UViG_3300009183_005911|3300009183|Ga0114974_10000105111 |
| APIS050 | 5.86E-51 | 1 | 97 | 2 | 112 | - | IMGVR_UViG_3300010885_038456|3300010885|Ga0133913_1000088583 |
| APIS074 | 8.65E-49 | 1 | 97 | 5 | 105 | - | IMGVR_UViG_3300042691_005779|3300042691|Ga0415385_0000706_88760_89182 |
| APIS082 | 7.60E-22 | 1 | 91 | 2 | 98 | - | MGV-GENOME-0365202_107 |
Family members, sources, and their hosts help
| APIS | Contig | Range |
|---|---|---|
| APIS035 | NC_047835.1 | 4114 - 8818 |
