Seed proteins |
Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS159 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | ||||||||||||||
| CLAN ID | CLAN003 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | ||||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | PF07275.16 | |||||||||||||
| Phrog | ||||||||||||||
| Host taxa | ;;;;;; | |||||||||||||
| Gene Location | Start: ; End: ; Strand: | |||||||||||||
| Description | ||||||||||||||
| Note | This family does not have seed protein, but we infer its function is defending restriction-modification (RM) because it belongs to the same clan as family APIS003,ardu which has seed protein. | |||||||||||||
Predicted 3D structure by alphafold2 with pTM = 0.72 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
Structure homologs against by afdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| AF-A0A132P918-F1-model_v4 | 37 | 207 | 1 | 166 | 0.8022 | 0.6994 |
| AF-A0A0H3GZQ3-F1-model_v4 | 32 | 207 | 1 | 168 | 0.7936 | 0.6967 |
| AF-A0A132ZEK5-F1-model_v4 | 37 | 207 | 1 | 167 | 0.797 | 0.6955 |
Structure homologs against by esmdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| MGYP003656451946.pdb | 16 | 207 | 5 | 194 | 0.859 | 0.8177 |
| MGYP003323835201.pdb | 41 | 207 | 1 | 170 | 0.907 | 0.8128 |
| MGYP001158282924.pdb | 21 | 207 | 1 | 177 | 0.8884 | 0.81 |
| MGYP001604904744.pdb | 41 | 207 | 1 | 170 | 0.888 | 0.7992 |
| MGYP001226369659.pdb | 41 | 207 | 1 | 170 | 0.8869 | 0.7964 |
| MGYP000444401356.pdb | 41 | 207 | 1 | 165 | 0.8972 | 0.7961 |
| MGYP000557273987.pdb | 19 | 207 | 1 | 181 | 0.8552 | 0.7855 |
| MGYP000526673471.pdb | 26 | 207 | 1 | 176 | 0.8668 | 0.7841 |
| MGYP003116609048.pdb | 37 | 207 | 4 | 172 | 0.8686 | 0.7823 |
| MGYP002642003215.pdb | 37 | 207 | 1 | 168 | 0.8771 | 0.7818 |
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS159 | IMGVR_UViG_3300040856_001282|3300040856|Ga0436647_0012686_635_1258 | 12 | HMM model | Member alignment |
|---|
Host Taxa distribution
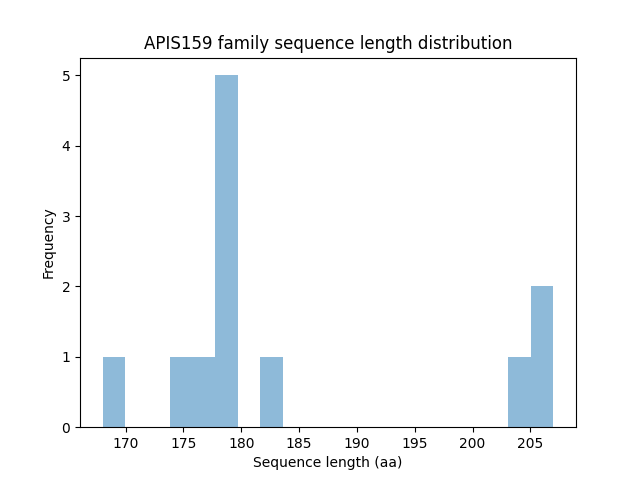
Length distribution