Seed proteins |
Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS133 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | CRISPR-Cas evasion by DNA repair | |||||||||||||
| CLAN ID | CLAN056 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | PMID:32556263 | |||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | PF03837.19 | |||||||||||||
| Phrog | phrog_12233,phrog_252,phrog_8375,phrog_10056 | |||||||||||||
| Host taxa | d__Bacteria;p__Proteobacteria;c__Gammaproteobacteria;o__Vibrionales;f__Vibrionaceae;g__Vibrio; | |||||||||||||
| Gene Location | Start: ; End: ; Strand: | |||||||||||||
| Description | CRISPR–Cas evasion by repairing double-strand DNA breaks via recombination between short sequence repeats | |||||||||||||
Predicted 3D structure by alphafold2 with pTM = 0.57 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
Structure homologs against by afdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| AF-Q32GM7-F1-model_v4 | 15 | 223 | 2 | 193 | 0.8259 | 0.5916 |
Structure homologs against by esmdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| MGYP001022669891.pdb | 20 | 211 | 1 | 187 | 0.9335 | 0.7162 |
| MGYP001049722605.pdb | 16 | 212 | 7 | 202 | 0.9236 | 0.7111 |
| MGYP000396468630.pdb | 30 | 219 | 1 | 173 | 0.9255 | 0.6945 |
| MGYP001042576686.pdb | 15 | 223 | 8 | 198 | 0.8818 | 0.6904 |
| MGYP001571145685.pdb | 6 | 203 | 1 | 170 | 0.9254 | 0.6878 |
| MGYP001563290713.pdb | 15 | 220 | 9 | 191 | 0.8884 | 0.6871 |
| MGYP003491535395.pdb | 16 | 218 | 10 | 194 | 0.8722 | 0.6823 |
| MGYP000196605117.pdb | 15 | 219 | 6 | 186 | 0.8824 | 0.6819 |
| MGYP000461655890.pdb | 15 | 191 | 1 | 176 | 0.9071 | 0.6808 |
| MGYP001010422329.pdb | 22 | 274 | 1 | 200 | 0.8652 | 0.6788 |
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS133 | YP_008997728.1 | 6 | HMM model | Member alignment |
|---|
Host Taxa distribution
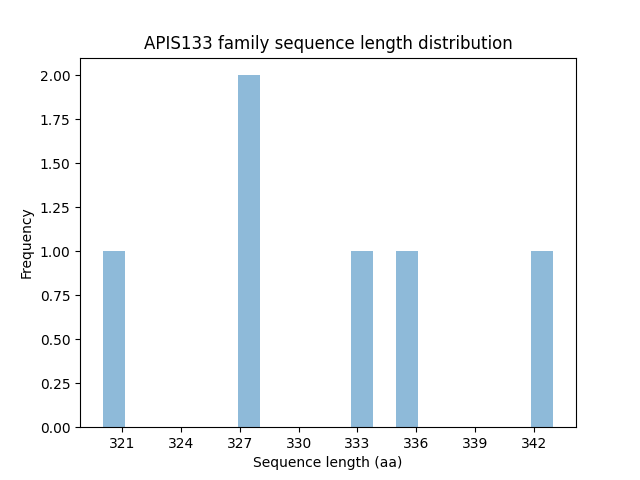
Length distribution