Seed proteins |
Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS060 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | ||||||||||||||
| CLAN ID | CLAN001 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | ||||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | PF00753.30 | |||||||||||||
| Phrog | phrog_392 | |||||||||||||
| Host taxa | ||||||||||||||
| Gene Location | Start: ; End: ; Strand: | |||||||||||||
| Description | ||||||||||||||
| Note | This family does not have seed protein, but we infer its function is defending pyrimidine cyclase system for antiphage resistance (Pycsar) because it belongs to the same clan as family APIS001 which has seed protein. | |||||||||||||
Predicted 3D structure by alphafold2 with pTM = 0.9 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
Structure homologs against by afdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| AF-A0A133CTF0-F1-model_v4 | 1 | 259 | 1 | 246 | 0.8428 | 0.7954 |
| AF-A0A0H3GHL8-F1-model_v4 | 5 | 263 | 1 | 244 | 0.8257 | 0.7871 |
| AF-O74545-F1-model_v4 | 1 | 261 | 6 | 301 | 0.8416 | 0.7757 |
| AF-P9WGC1-F1-model_v4 | 1 | 262 | 18 | 272 | 0.8097 | 0.7739 |
| AF-Q58769-F1-model_v4 | 1 | 263 | 3 | 260 | 0.7998 | 0.7715 |
| AF-I1KRW8-F1-model_v4 | 1 | 261 | 3 | 311 | 0.8437 | 0.7661 |
| AF-I1K3H5-F1-model_v4 | 1 | 261 | 3 | 308 | 0.8375 | 0.7639 |
| AF-A0A1P8B5J8-F1-model_v4 | 1 | 261 | 11 | 318 | 0.8486 | 0.7637 |
| AF-P50474-F1-model_v4 | 1 | 263 | 27 | 282 | 0.8106 | 0.7615 |
| AF-A0A0P0W0M3-F1-model_v4 | 1 | 261 | 5 | 312 | 0.8387 | 0.7603 |
Structure homologs against by esmdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| MGYP002654867125.pdb | 1 | 259 | 3 | 255 | 0.9415 | 0.9194 |
| MGYP000856702051.pdb | 1 | 260 | 3 | 268 | 0.8858 | 0.8648 |
| MGYP000228473758.pdb | 1 | 261 | 1 | 255 | 0.8908 | 0.8634 |
| MGYP001465423269.pdb | 1 | 259 | 1 | 253 | 0.8918 | 0.8615 |
| MGYP003624123598.pdb | 1 | 260 | 1 | 259 | 0.8802 | 0.8609 |
| MGYP003194564429.pdb | 1 | 259 | 1 | 252 | 0.8939 | 0.8594 |
| MGYP001278173685.pdb | 1 | 259 | 1 | 252 | 0.8927 | 0.8585 |
| MGYP003452926687.pdb | 1 | 260 | 2 | 266 | 0.8755 | 0.8581 |
| MGYP003606950373.pdb | 1 | 261 | 4 | 256 | 0.8816 | 0.8564 |
| MGYP003952926961.pdb | 1 | 260 | 2 | 254 | 0.8857 | 0.8558 |
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS060 | IMGVR_UViG_3300024258_000005|3300024258|Ga0233440_1000037199 | 3 | HMM model | Member alignment |
|---|
Host Taxa distribution
Length distribution
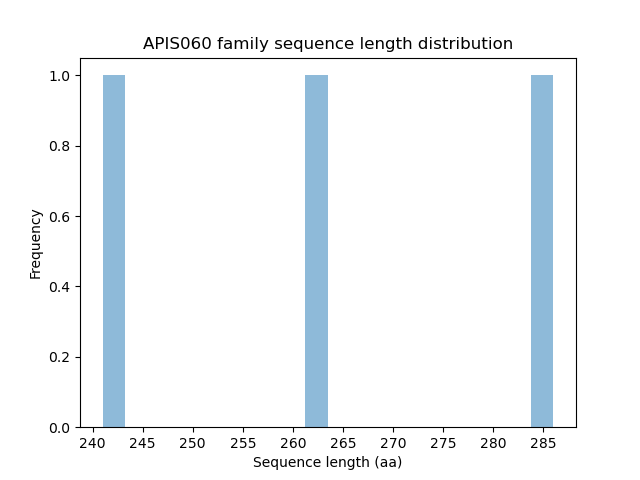
Homology to other families download full data without filtering help
Clan ID: CLAN001
| Hit APIS family | E-Value | Query Start | Query End | Hit Start | Hit End | Immune system | Seed protein |
|---|---|---|---|---|---|---|---|
| APIS058 | 2.87E-77 | 1 | 262 | 16 | 271 | - | IMGVR_UViG_3300022562_000702|3300022562|Ga0247387_1000200120 |
| APIS054 | 8.02E-55 | 1 | 263 | 3 | 266 | - | IMGVR_UViG_3300014204_000482|3300014204|Ga0172381_1000105623 |
| APIS066 | 8.92E-49 | 1 | 262 | 1 | 296 | - | IMGVR_UViG_3300035396_000446|3300035396|Ga0393109_006065_59543_60412 |
| APIS073 | 2.51E-41 | 1 | 262 | 23 | 288 | - | IMGVR_UViG_3300042197_004918|3300042197|Ga0451494_0000046_37885_38742 |
| APIS064 | 7.55E-41 | 1 | 259 | 1 | 258 | - | IMGVR_UViG_3300033145_003233|3300033145|Ga0366840_1000004654 |
| APIS062 | 7.97E-40 | 1 | 232 | 1 | 243 | - | IMGVR_UViG_3300028374_000006|3300028374|Ga0306906_100003345 |
| APIS001 | 9.98E-39 | 1 | 260 | 20 | 271 | pyrimidine cyclase system for antiphage resistance (Pycsar) | YP_009840594.1 |
| APIS076 | 1.72E-37 | 2 | 261 | 7 | 261 | - | IMGVR_UViG_3300044693_000096|3300044693|Ga0466961_0000159_42271_43065 |
| APIS044 | 2.11E-24 | 2 | 248 | 1 | 233 | - | HM144387_00160 |
| APIS041 | 1.18E-22 | 1 | 257 | 2 | 247 | - | IMGVR_UViG_2832702071_000001|2832702071|2832704097 |
Family members, sources, and their hosts help
| Member | pident | evalue | bitscore | qcovs | phage | host | SourceDB | Pfam | Phrog |
|---|---|---|---|---|---|---|---|---|---|
| IMGVR_UViG_3300013795_000141|3300013795|Ga0180408_1001003 | 42.593 | 8.76e-55 | 185 | 99% | IMGVR | PF00753 | phrog_392 | ||
| IMGVR_UViG_3300024258_000005|3300024258|Ga0233440_1000037199 | 100.000 | 0.0 | 542 | 100% | IMGVR | PF00753 | phrog_392 | ||
| IMGVR_UViG_3300035188_001312|3300035188|Ga0335059_0000026_130956_131816 | 41.219 | 2.02e-60 | 201 | 97% | IMGVR | PF12706 | phrog_392 |
