Seed proteins |
Seed genomic context | Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS040 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | ||||||||||||||
| CLAN ID | CLAN030 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | ||||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | PF14354.9 | |||||||||||||
| Phrog | phrog_6422,phrog_599 | |||||||||||||
| Host taxa | d__Bacteria;p__Proteobacteria;c__Gammaproteobacteria;o__Enterobacterales;f__Enterobacteriaceae;g__Escherichia;s__Escherichia coli | |||||||||||||
| Gene Location | Start: 3768; End: 4196; Strand: - | |||||||||||||
| Description | ||||||||||||||
| Note | This family does not have seed protein, but we infer its function is defending restriction-modification (RM) because it belongs to the same clan as family APIS033 which has seed protein. | |||||||||||||
Seed genomic context Download table help
| Genome accession | Start | End | Strand | Protein ID | Length | Molecular weight | Charge | Isoelectric point | Function | Phrog | PFAM |
|---|---|---|---|---|---|---|---|---|---|---|---|
| IMGVR_UViG_2778261704_000001 | 1 | 618 | - | IMGVR_UViG_2778261704_000001|2778261704|2781089220 | 206 | 22937.34 | 4.5 | 9.7270 | phage antirepressor YoqD-like protein | phrog_5689 ; phrog_15709 ; phrog_9438 ; phrog_4936 ; phrog_15968 ; phrog_63 ; phrog_10324 | PF03374.17 |
| IMGVR_UViG_2778261704_000001 | 1 | 618 | - | IMGVR_UViG_2778261704_000001|2778261704|2781089220 | 206 | 22937.34 | 4.5 | 9.7270 | phage antirepressor YoqD-like protein | phrog_5689 ; phrog_15709 ; phrog_9438 ; phrog_4936 ; phrog_15968 ; phrog_63 ; phrog_10324 | PF09669.13 |
| IMGVR_UViG_2778261704_000001 | 693 | 1373 | - | IMGVR_UViG_2778261704_000001|2778261704|2781089221 | 226 | 26347.07 | 6.0 | 8.5952 | phage anti-repressor protein | phrog_7755 ; phrog_4936 ; phrog_2812 ; phrog_8244 ; phrog_4792 ; phrog_2379 ; phrog_15086 ; phrog_2127 ; phrog_1760 ; phrog_377 ; phrog_22471 ; phrog_5060 ; phrog_1726 | PF10543.12 |
| IMGVR_UViG_2778261704_000001 | 693 | 1373 | - | IMGVR_UViG_2778261704_000001|2778261704|2781089221 | 226 | 26347.07 | 6.0 | 8.5952 | phage anti-repressor protein | phrog_7755 ; phrog_4936 ; phrog_2812 ; phrog_8244 ; phrog_4792 ; phrog_2379 ; phrog_15086 ; phrog_2127 ; phrog_1760 ; phrog_377 ; phrog_22471 ; phrog_5060 ; phrog_1726 | PF10548.12 |
| IMGVR_UViG_2778261704_000001 | 1629 | 2387 | - | IMGVR_UViG_2778261704_000001|2778261704|2781089222 | 252 | 29143.28 | 8.5 | 9.5155 | ORF6N domain-containing protein | phrog_463 ; phrog_7755 ; phrog_4936 ; phrog_450 ; phrog_995 ; phrog_24476 ; phrog_8244 ; phrog_4792 ; phrog_2379 ; phrog_15086 ; phrog_2127 ; phrog_1760 ; phrog_377 ; phrog_22471 ; phrog_17980 ; phrog_1726 | PF10543.12 |
| IMGVR_UViG_2778261704_000001 | 2440 | 2523 | - | IMGVR_UViG_2778261704_000001|2778261704|2781089223 | (miscRNA ) | ||||||
| IMGVR_UViG_2778261704_000001 | 2841 | 3368 | - | IMGVR_UViG_2778261704_000001|2778261704|2781089224 | 175 | 19737.60 | -1.5 | 5.6965 | phage N-6-adenine-methyltransferase | phrog_6083 ; phrog_11720 ; phrog_111 ; phrog_33262 ; phrog_22740 | PF05869.14 |
| IMGVR_UViG_2778261704_000001 | 3365 | 3811 | - | IMGVR_UViG_2778261704_000001|2778261704|2781089225 | 148 | 17363.98 | 8.5 | 10.6047 | NinB protein | phrog_72 | PF05772.15 |
| IMGVR_UViG_2778261704_000001 | 3768 | 4196 | - | IMGVR_UViG_2778261704_000001|2778261704|2781089226 | 142 | 15541.78 | 0.5 | 6.6684 | endodeoxyribonuclease RalR | phrog_6422 ; phrog_1603 ; phrog_599 | PF14354.9 |
Seed genomic context- JBrowse
Predicted 3D structure by alphafold2 with pTM = 0.42 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
No homologs found in AlphaFold database
No homologs found in esmfold database
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS040 | IMGVR_UViG_2778261704_000001|2778261704|2781089226 | 8 | HMM model | Member alignment |
|---|
Host Taxa distribution
Length distribution
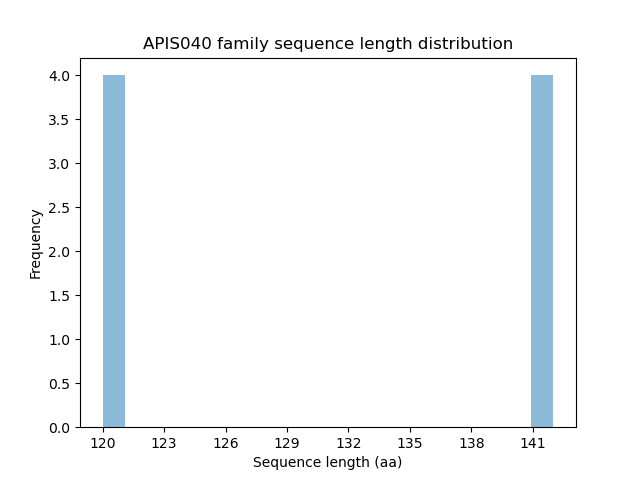
Homology to other families download full data without filtering help
Clan ID: CLAN030
| Hit APIS family | E-Value | Query Start | Query End | Hit Start | Hit End | Immune system | Seed protein |
|---|---|---|---|---|---|---|---|
| APIS033 | 7.99E-52 | 63 | 142 | 1 | 80 | restriction-modification (RM) | YP_009168126.1 |
