Seed proteins |
Seed genomic context | Seed protein 3D structures |
Structural homologs |
Family infomation |
Homology to other families |
Family members, sources, and their hosts
Seed protein information help
| APIS family ID | APIS029 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Inhibited defense system | Thoeris | |||||||||||||
| CLAN ID | CLAN023 | |||||||||||||
| Seed protein property |
|
|||||||||||||
| Phage property |
|
|||||||||||||
| PMID/References | Yirmiya et al., 2023 | |||||||||||||
| PDB structures | ; | |||||||||||||
| Pfam domains | PF11195.11 | |||||||||||||
| Phrog | phrog_2709,phrog_30130,phrog_872,phrog_1048 | |||||||||||||
| Host taxa | d__Bacteria;p__Firmicutes;c__Bacilli;o__Bacillales;f__Bacillaceae;g__Bacillus; | |||||||||||||
| Gene Location | Start: 127121; End: 127390; Strand: + | |||||||||||||
| Description | Tad2 is a "sponge" that sequesters the immune signaling molecules produced by Thoeris TIR-domain proteins in response to phage. | |||||||||||||
Seed genomic context Download table help
| Genome accession | Start | End | Strand | Protein ID | Length | Molecular weight | Charge | Isoelectric point | Function | Phrog | PFAM |
|---|---|---|---|---|---|---|---|---|---|---|---|
| NC_011421.1 | 125619 | 125978 | + | YP_002300459.1 | 119 | 13655.34 | -4.5 | 4.6850 | gp34.56 | phrog_27569 | |
| NC_011421.1 | 125978 | 126109 | + | YP_002300460.1 | 43 | 5007.66 | -8.5 | 3.7350 | gp34.61 | ||
| NC_011421.1 | 126099 | 126452 | + | YP_002300461.1 | 117 | 13701.68 | -1.5 | 5.7247 | gp34.62 | phrog_26195 | |
| NC_011421.1 | 126433 | 126711 | + | YP_002300462.1 | 92 | 11143.87 | 8.0 | 10.1732 | gp34.63 | phrog_18481 | |
| NC_011421.1 | 126711 | 127121 | + | YP_002300463.1 | 136 | 15653.55 | 4.0 | 9.6717 | gp34.64 | phrog_18481 | |
| NC_011421.1 | 127121 | 127390 | + | YP_002300464.1 | 89 | 10029.45 | -2.5 | 4.6684 | gp34.65 | phrog_1048 ; phrog_2709 ; phrog_30130 ; phrog_872 | PF11195.11 |
| NC_011421.1 | 127418 | 127645 | + | YP_002300465.1 | 75 | 8736.63 | 5.5 | 9.4374 | gp34.66 | phrog_24625 | |
| NC_011421.1 | 127647 | 127835 | + | YP_002300466.1 | 62 | 7099.01 | -7.5 | 3.9326 | gp34.71 | phrog_31357 | |
| NC_011421.1 | 127851 | 128120 | + | YP_002300467.1 | 89 | 10521.74 | -7.0 | 4.4526 | gp34.72 | phrog_7040 | |
| NC_011421.1 | 128117 | 128257 | + | YP_002300468.1 | 46 | 5081.72 | -5.5 | 4.1087 | gp34.73 | phrog_20907 ; phrog_22847 | |
| NC_011421.1 | 128282 | 128593 | + | YP_002300469.1 | 103 | 11516.85 | -8.0 | 4.3141 | gp34.74 | phrog_25367 |
Seed genomic context- JBrowse
Predicted 3D structure by alphafold2 with pTM = 0.79 Download help
pTM is for the estimate of the TM-score, which is obtained from a pairwise error prediction. The higher pTM score indicates better model quality.
pLDDT is for per-residue accuracy of the structure, which representes the quality of the residue. A higher value indicates better prediction accuracy. More detail please see AlphaFold .
Residues were colored according to plddt ( blue-> high quality; red-> low quality ).
Full Sequence
90 < plddt <=100 ;
70 < plddt <= 90 ;
50 < plddt <= 70 ;
0 <= plddt <= 50 ;
Download
help
Structural homologs
Structure homologs against by afdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| AF-A0A132Z7R2-F1-model_v4 | 5 | 89 | 1 | 73 | 0.8947 | 0.7892 |
Structure homologs against by esmdb (html) ; download full result table
| Hit | Query start | Query end | Hit start | Hit end | TMscore(ali) | TMscore(avg) |
|---|---|---|---|---|---|---|
| MGYP001070471033.pdb | 1 | 89 | 1 | 89 | 0.8895 | 0.8672 |
| MGYP003762584705.pdb | 1 | 89 | 1 | 76 | 0.9272 | 0.8422 |
| MGYP001563495202.pdb | 5 | 88 | 1 | 69 | 0.9603 | 0.842 |
| MGYP003431837514.pdb | 5 | 89 | 1 | 79 | 0.8928 | 0.8273 |
| MGYP003457190342.pdb | 1 | 89 | 1 | 76 | 0.9066 | 0.8247 |
| MGYP001121567902.pdb | 5 | 89 | 1 | 86 | 0.8679 | 0.8246 |
| MGYP000961172731.pdb | 1 | 88 | 1 | 75 | 0.9111 | 0.8246 |
| MGYP000057650656.pdb | 5 | 89 | 1 | 72 | 0.924 | 0.8218 |
| MGYP000184816965.pdb | 1 | 89 | 1 | 77 | 0.9055 | 0.8216 |
| MGYP000920153130.pdb | 3 | 89 | 1 | 74 | 0.9065 | 0.8208 |
Family information Download help
| Fam ID | Seed protein | Member count | Model | Alignment | APIS029 | YP_002300464.1 | 132 | HMM model | Member alignment |
|---|
Host Taxa distribution
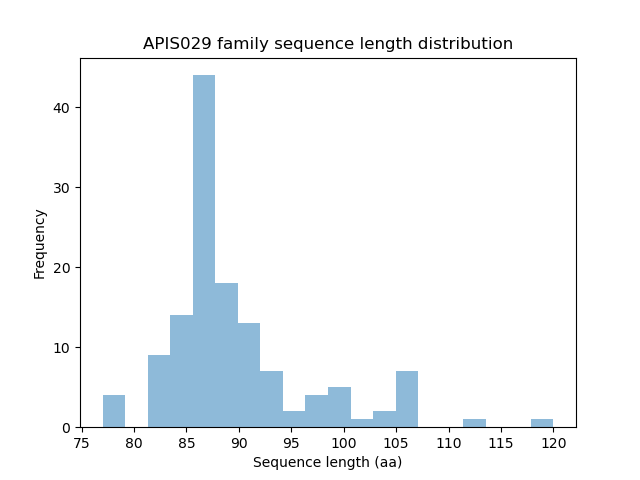
Length distribution
Homology to other families download full data without filtering help
Clan ID: CLAN023
| Hit APIS family | E-Value | Query Start | Query End | Hit Start | Hit End | Immune system | Seed protein |
|---|---|---|---|---|---|---|---|
| APIS079 | 4.32E-48 | 1 | 85 | 88 | 171 | - | MF042361_00008 |
| APIS052 | 2.34E-40 | 1 | 85 | 77 | 162 | - | IMGVR_UViG_3300012991_000489|3300012991|Ga0157148_10003866 |
| APIS043 | 2.82E-40 | 1 | 85 | 122 | 208 | - | MH238467_00052 |
| APIS045 | 2.91E-34 | 2 | 85 | 24 | 115 | - | IMGVR_UViG_3300005805_000580|3300005805|Ga0079957_100145420 |
Family members, sources, and their hosts help
| APIS | Contig | Range |
|---|---|---|
| APIS029 | NC_011421.1 | 125619 - 128593 |
