You are browsing environment: FUNGIDB
CAZyme Information: jhhlp_008289-t41_1-p1
You are here: Home > Sequence: jhhlp_008289-t41_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Lomentospora prolificans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Microascaceae; Lomentospora; Lomentospora prolificans | |||||||||||
| CAZyme ID | jhhlp_008289-t41_1-p1 | |||||||||||
| CAZy Family | GT31 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE5 | 48 | 221 | 1.9e-35 | 0.9788359788359788 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395860 | Cutinase | 7.52e-30 | 48 | 221 | 2 | 171 | Cutinase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.26e-109 | 8 | 223 | 6 | 230 | |
| 1.67e-66 | 23 | 225 | 36 | 242 | |
| 1.66e-39 | 78 | 225 | 1 | 149 | |
| 9.17e-33 | 46 | 206 | 29 | 198 | |
| 3.98e-31 | 46 | 206 | 29 | 198 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.84e-19 | 48 | 220 | 79 | 244 | Structure of cutinase from Trichoderma reesei in its native form. [Trichoderma reesei QM6a],4PSD_A Structure of Trichoderma reesei cutinase native form. [Trichoderma reesei QM6a],4PSE_A Trichoderma reesei cutinase in complex with a C11Y4 phosphonate inhibitor [Trichoderma reesei QM6a],4PSE_B Trichoderma reesei cutinase in complex with a C11Y4 phosphonate inhibitor [Trichoderma reesei QM6a] |
|
| 1.38e-14 | 48 | 206 | 23 | 183 | Chain A, cutinase [Malbranchea cinnamomea] |
|
| 2.57e-13 | 46 | 206 | 1 | 178 | Chain A, Cutinase-like enzyme [Moesziomyces antarcticus],7CC4_B Chain B, Cutinase-like enzyme [Moesziomyces antarcticus],7CW1_A Chain A, Cutinase-like enzyme [Moesziomyces antarcticus],7CW1_B Chain B, Cutinase-like enzyme [Moesziomyces antarcticus] |
|
| 1.79e-10 | 48 | 194 | 6 | 169 | A novel cutinase-like protein from Cryptococcus sp. [Cryptococcus sp. S-2],2CZQ_B A novel cutinase-like protein from Cryptococcus sp. [Cryptococcus sp. S-2] |
|
| 2.12e-10 | 48 | 206 | 17 | 176 | Humicola insolens cutinase [Humicola insolens],4OYY_B Humicola insolens cutinase [Humicola insolens],4OYY_C Humicola insolens cutinase [Humicola insolens],4OYY_D Humicola insolens cutinase [Humicola insolens],4OYY_E Humicola insolens cutinase [Humicola insolens],4OYY_F Humicola insolens cutinase [Humicola insolens],4OYY_G Humicola insolens cutinase [Humicola insolens],4OYY_H Humicola insolens cutinase [Humicola insolens],4OYY_I Humicola insolens cutinase [Humicola insolens],4OYY_J Humicola insolens cutinase [Humicola insolens],4OYY_K Humicola insolens cutinase [Humicola insolens],4OYY_L Humicola insolens cutinase [Humicola insolens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.14e-19 | 48 | 219 | 79 | 243 | Cutinase cut1 OS=Trichoderma harzianum OX=5544 GN=cut1 PE=1 SV=1 |
|
| 8.54e-19 | 48 | 220 | 79 | 244 | Cutinase OS=Hypocrea jecorina (strain QM6a) OX=431241 GN=TRIREDRAFT_60489 PE=3 SV=2 |
|
| 8.54e-19 | 48 | 220 | 79 | 244 | Cutinase OS=Hypocrea jecorina (strain ATCC 56765 / BCRC 32924 / NRRL 11460 / Rut C-30) OX=1344414 GN=M419DRAFT_76732 PE=1 SV=1 |
|
| 4.74e-16 | 48 | 220 | 38 | 210 | Cutinase 1 OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=cut1 PE=2 SV=1 |
|
| 8.85e-16 | 13 | 206 | 5 | 196 | Probable cutinase 1 OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=NFIA_084890 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
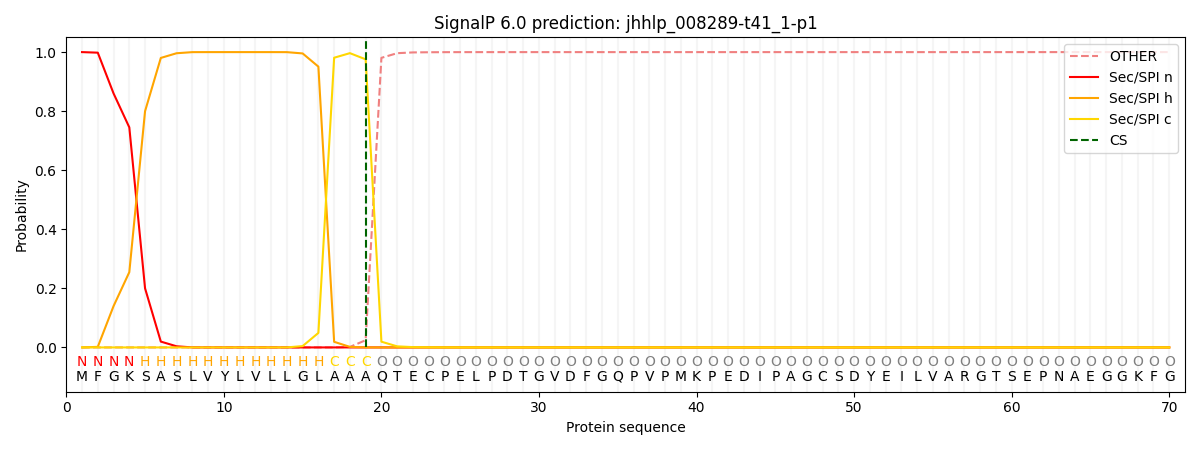
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000221 | 0.999744 | CS pos: 19-20. Pr: 0.9755 |
