You are browsing environment: FUNGIDB
CAZyme Information: jhhlp_003374-t41_1-p1
You are here: Home > Sequence: jhhlp_003374-t41_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Lomentospora prolificans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Microascaceae; Lomentospora; Lomentospora prolificans | |||||||||||
| CAZyme ID | jhhlp_003374-t41_1-p1 | |||||||||||
| CAZy Family | CE5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.4:5 | 3.2.1.151:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH12 | 98 | 245 | 3.3e-42 | 0.9871794871794872 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396303 | Glyco_hydro_12 | 1.59e-44 | 40 | 246 | 1 | 207 | Glycosyl hydrolase family 12. |
| 235746 | PRK06215 | 2.36e-20 | 2 | 192 | 12 | 184 | hypothetical protein; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 8.62e-114 | 12 | 246 | 13 | 248 | |
| 1.95e-103 | 1 | 245 | 1 | 248 | |
| 2.77e-103 | 1 | 245 | 1 | 248 | |
| 2.77e-103 | 1 | 245 | 1 | 248 | |
| 2.77e-103 | 1 | 245 | 1 | 248 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.43e-89 | 19 | 245 | 16 | 242 | Crystal Structure of the Family 12 Xyloglucanase from Aspergillus niveus [Aspergillus niveus],4NPR_B Crystal Structure of the Family 12 Xyloglucanase from Aspergillus niveus [Aspergillus niveus] |
|
| 3.16e-79 | 25 | 246 | 9 | 229 | Crystal structure of XEG [Aspergillus aculeatus],3VL9_A Crystal structure of xeg-xyloglucan [Aspergillus aculeatus],3VL9_B Crystal structure of xeg-xyloglucan [Aspergillus aculeatus] |
|
| 2.90e-78 | 31 | 246 | 8 | 222 | Crystal structure of xeg-edgp [Aspergillus aculeatus],3VLB_D Crystal structure of xeg-edgp [Aspergillus aculeatus] |
|
| 3.00e-51 | 31 | 246 | 4 | 219 | Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 [Aspergillus aculeatus],5GM3_B Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 [Aspergillus aculeatus] |
|
| 2.40e-50 | 31 | 246 | 4 | 219 | Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus],5GM4_B Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus],5GM4_C Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus],5GM4_D Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus],5GM4_E Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus],5GM4_F Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus],5GM4_G Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellotetrose [Aspergillus aculeatus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.71e-89 | 13 | 245 | 8 | 237 | Probable xyloglucan-specific endo-beta-1,4-glucanase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=xgeA PE=3 SV=1 |
|
| 3.71e-89 | 13 | 245 | 8 | 237 | Probable xyloglucan-specific endo-beta-1,4-glucanase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=xgeA PE=3 SV=1 |
|
| 7.45e-89 | 13 | 245 | 8 | 237 | Probable xyloglucan-specific endo-beta-1,4-glucanase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=xgeA PE=3 SV=1 |
|
| 3.97e-79 | 26 | 245 | 29 | 249 | Probable xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=xgeA PE=3 SV=2 |
|
| 5.95e-79 | 20 | 246 | 14 | 241 | Probable xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=xgeA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
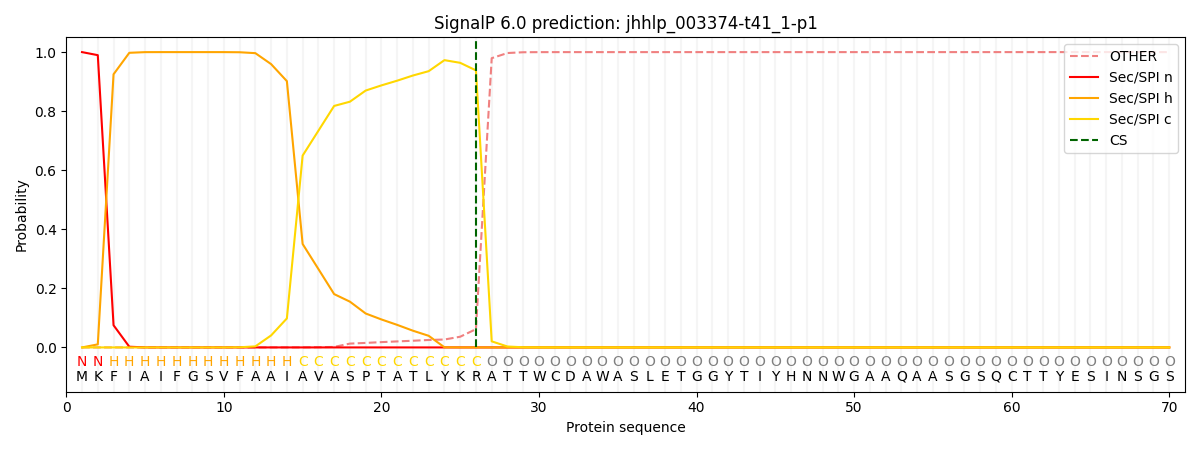
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000239 | 0.999731 | CS pos: 26-27. Pr: 0.9381 |
