You are browsing environment: FUNGIDB
CAZyme Information: jhhlp_003285-t41_1-p1
You are here: Home > Sequence: jhhlp_003285-t41_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
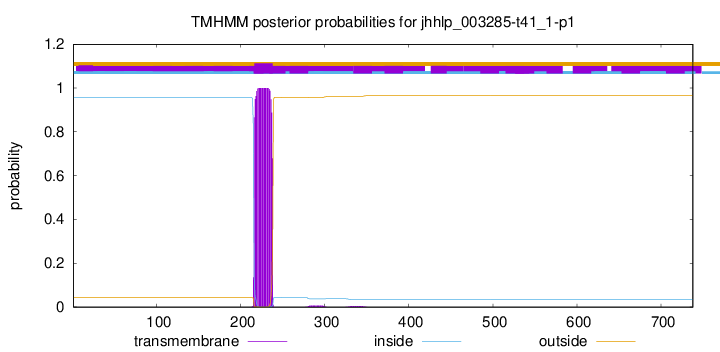
TMHMM annotations
Basic Information help
| Species | Lomentospora prolificans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Microascaceae; Lomentospora; Lomentospora prolificans | |||||||||||
| CAZyme ID | jhhlp_003285-t41_1-p1 | |||||||||||
| CAZy Family | CE4 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.58:2 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 374 | 697 | 6.2e-107 | 0.9966996699669967 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 225344 | BglC | 1.19e-48 | 320 | 714 | 12 | 382 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| 395098 | Cellulase | 3.18e-06 | 379 | 562 | 24 | 195 | Cellulase (glycosyl hydrolase family 5). |
| 397943 | Radical_SAM | 0.001 | 377 | 445 | 83 | 150 | Radical SAM superfamily. Radical SAM proteins catalyze diverse reactions, including unusual methylations, isomerisation, sulphur insertion, ring formation, anaerobic oxidation and protein radical formation. |
| 236766 | rne | 0.006 | 89 | 142 | 599 | 649 | ribonuclease E; Reviewed |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.67e-299 | 156 | 738 | 130 | 711 | |
| 3.35e-286 | 147 | 731 | 91 | 682 | |
| 8.37e-282 | 158 | 731 | 103 | 682 | |
| 2.67e-279 | 164 | 737 | 109 | 695 | |
| 2.67e-279 | 164 | 737 | 109 | 695 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.80e-68 | 315 | 722 | 3 | 399 | Crystal structure of exo-1,3-beta glucanse from Saccharomyces cerevisiae [Saccharomyces cerevisiae],1H4P_B Crystal structure of exo-1,3-beta glucanse from Saccharomyces cerevisiae [Saccharomyces cerevisiae] |
|
| 3.16e-66 | 321 | 722 | 4 | 387 | Exo-b-(1,3)-glucanase From Candida Albicans [Candida albicans] |
|
| 6.12e-66 | 321 | 722 | 4 | 387 | Exo-b-(1,3)-glucanase From Candida Albicans At 1.85 A Resolution [Candida albicans],1EQC_A Exo-b-(1,3)-glucanase From Candida Albicans In Complex With Castanospermine At 1.85 A [Candida albicans] |
|
| 7.21e-66 | 321 | 722 | 10 | 393 | Exo-B-(1,3)-Glucanase from Candida Albicans in complex with unhydrolysed and covalently linked 2,4-dinitrophenyl-2-deoxy-2-fluoro-B-D-glucopyranoside at 1.9 A [Candida albicans] |
|
| 1.36e-65 | 321 | 722 | 9 | 392 | F144Y/F258Y Double Mutant of Exo-beta-1,3-glucanase from Candida albicans at 2 A [Candida albicans] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.34e-212 | 158 | 738 | 239 | 833 | Probable glucan 1,3-beta-glucosidase D OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=exgD PE=3 SV=1 |
|
| 1.34e-212 | 158 | 738 | 239 | 833 | Probable glucan 1,3-beta-glucosidase D OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=exgD PE=3 SV=1 |
|
| 1.25e-210 | 158 | 738 | 240 | 834 | Probable glucan 1,3-beta-glucosidase D OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=exgD PE=3 SV=1 |
|
| 2.27e-210 | 250 | 738 | 345 | 831 | Probable glucan 1,3-beta-glucosidase D OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=exgD PE=3 SV=2 |
|
| 2.27e-210 | 250 | 738 | 345 | 831 | Probable glucan 1,3-beta-glucosidase D OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=exgD PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
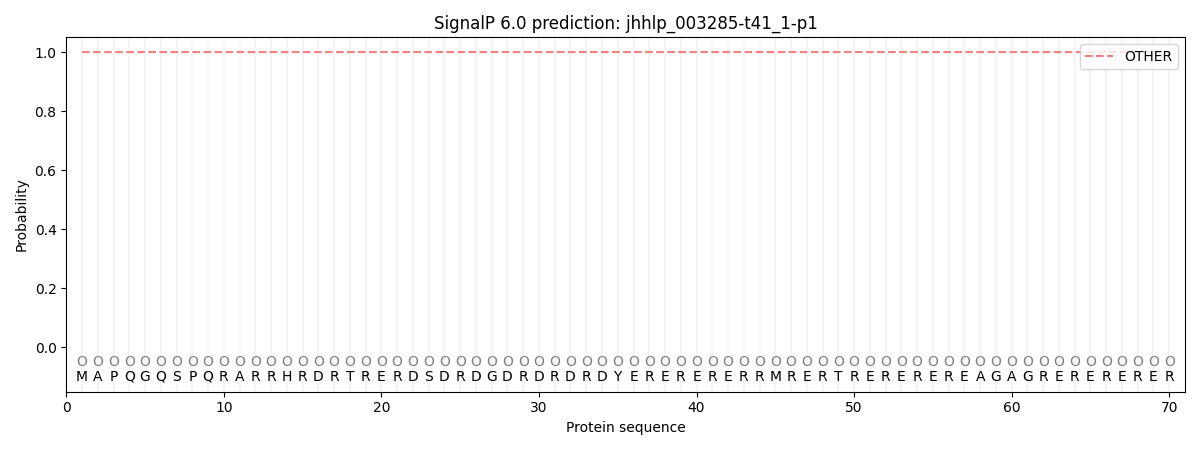
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000062 | 0.000000 |