You are browsing environment: FUNGIDB
CAZyme Information: jhhlp_002964-t41_1-p1
You are here: Home > Sequence: jhhlp_002964-t41_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Lomentospora prolificans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Microascaceae; Lomentospora; Lomentospora prolificans | |||||||||||
| CAZyme ID | jhhlp_002964-t41_1-p1 | |||||||||||
| CAZy Family | CE1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA3 | 49 | 662 | 2.1e-144 | 0.9947183098591549 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 235000 | PRK02106 | 6.32e-65 | 49 | 664 | 5 | 535 | choline dehydrogenase; Validated |
| 225186 | BetA | 5.45e-59 | 49 | 665 | 7 | 538 | Choline dehydrogenase or related flavoprotein [Lipid transport and metabolism, General function prediction only]. |
| 274888 | Rv0697 | 6.94e-43 | 51 | 660 | 2 | 486 | dehydrogenase, Rv0697 family. This model describes a set of dehydrogenases belonging to the glucose-methanol-choline oxidoreductase (GMC oxidoreductase) family. Members of the present family are restricted to Actinobacterial genome contexts containing also members of families TIGR03962 and TIGR03969 (the mycofactocin system), and are proposed to be uniform in function. |
| 366272 | GMC_oxred_N | 7.87e-28 | 118 | 382 | 12 | 217 | GMC oxidoreductase. This family of proteins bind FAD as a cofactor. |
| 398739 | GMC_oxred_C | 3.69e-24 | 509 | 656 | 3 | 143 | GMC oxidoreductase. This domain found associated with pfam00732. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.36e-150 | 50 | 666 | 25 | 607 | |
| 9.84e-149 | 45 | 661 | 34 | 628 | |
| 1.66e-146 | 45 | 661 | 35 | 635 | |
| 5.54e-145 | 24 | 663 | 18 | 634 | |
| 3.51e-143 | 35 | 663 | 31 | 646 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.36e-117 | 13 | 668 | 2 | 644 | Structural insights on a new fungal aryl-alcohol oxidase [Thermothelomyces thermophilus],6O9N_B Structural insights on a new fungal aryl-alcohol oxidase [Thermothelomyces thermophilus] |
|
| 2.70e-90 | 46 | 662 | 2 | 584 | Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE2_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE3_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE4_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE4_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE5_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE5_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE6_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE6_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE7_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE7_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],7AA2_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],7AA2_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495] |
|
| 2.47e-52 | 19 | 662 | 12 | 599 | Crystal structure of pyranose dehydrogenase from Agaricus meleagris, wildtype [Leucoagaricus meleagris] |
|
| 6.41e-51 | 50 | 661 | 6 | 565 | Crystal structure of Aspergillus flavus FAD glucose dehydrogenase [Aspergillus flavus NRRL3357],4YNU_A Crystal structure of Aspergillus flavus FADGDH in complex with D-glucono-1,5-lactone [Aspergillus flavus NRRL3357] |
|
| 1.39e-44 | 50 | 661 | 17 | 588 | Chain A, Oligosaccharide dehydrogenase [Trametes cinnabarina],6XUU_A Chain A, Oligosaccharide dehydrogenase [Trametes cinnabarina],6XUV_A Chain A, Oligosaccharide dehydrogenase [Trametes cinnabarina] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.23e-92 | 39 | 667 | 27 | 620 | Dehydrogenase xptC OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=xptC PE=3 SV=1 |
|
| 5.11e-77 | 51 | 662 | 41 | 631 | Dehydrogenase ARMGADRAFT_1018426 OS=Armillaria gallica OX=47427 GN=ARMGADRAFT_1018426 PE=1 SV=1 |
|
| 2.20e-65 | 50 | 660 | 47 | 615 | GMC oxidoreductase family protein Mala s 12.0101 OS=Malassezia sympodialis OX=76777 PE=1 SV=2 |
|
| 2.20e-65 | 50 | 660 | 47 | 615 | GMC oxidoreductase family protein Mala s 12 OS=Malassezia sympodialis (strain ATCC 42132) OX=1230383 GN=MSY001_2108 PE=1 SV=1 |
|
| 6.08e-64 | 38 | 666 | 2 | 614 | Dehydrogenase mpl7 OS=Monascus purpureus OX=5098 GN=mpl7 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
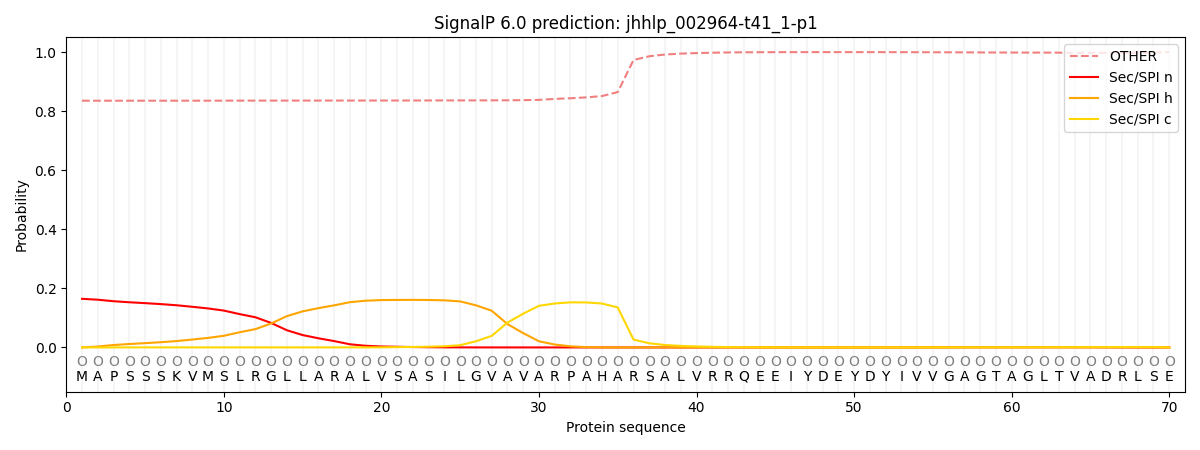
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.842406 | 0.157587 |
