You are browsing environment: FUNGIDB
CAZyme Information: evm.model.AC2VRR_s00168g42-p1
You are here: Home > Sequence: evm.model.AC2VRR_s00168g42-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Albugo candida | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Albuginaceae; Albugo; Albugo candida | |||||||||||
| CAZyme ID | evm.model.AC2VRR_s00168g42-p1 | |||||||||||
| CAZy Family | GH72 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT8 | 57 | 263 | 3.6e-32 | 0.8132295719844358 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 133018 | GT8_Glycogenin | 6.56e-52 | 44 | 320 | 1 | 240 | Glycogenin belongs the GT 8 family and initiates the biosynthesis of glycogen. Glycogenin initiates the biosynthesis of glycogen by incorporating glucose residues through a self-glucosylation reaction at a Tyr residue, and then acts as substrate for chain elongation by glycogen synthase and branching enzyme. It contains a conserved DxD motif and an N-terminal beta-alpha-beta Rossmann-like fold that are common to the nucleotide-binding domains of most glycosyltransferases. The DxD motif is essential for coordination of the catalytic divalent cation, most commonly Mn2+. Glycogenin can be classified as a retaining glycosyltransferase, based on the relative anomeric stereochemistry of the substrate and product in the reaction catalyzed. It is placed in glycosyltransferase family 8 which includes lipopolysaccharide glucose and galactose transferases and galactinol synthases. |
| 215090 | PLN00176 | 7.00e-12 | 45 | 208 | 24 | 219 | galactinol synthase |
| 132996 | Glyco_transf_8 | 1.70e-11 | 57 | 229 | 9 | 213 | Members of glycosyltransferase family 8 (GT-8) are involved in lipopolysaccharide biosynthesis and glycogen synthesis. Members of this family are involved in lipopolysaccharide biosynthesis and glycogen synthesis. GT-8 comprises enzymes with a number of known activities: lipopolysaccharide galactosyltransferase, lipopolysaccharide glucosyltransferase 1, glycogenin glucosyltransferase, and N-acetylglucosaminyltransferase. GT-8 enzymes contains a conserved DXD motif which is essential in the coordination of a catalytic divalent cation, most commonly Mn2+. |
| 133064 | GT8_GNT1 | 2.79e-11 | 44 | 299 | 1 | 238 | GNT1 is a fungal enzyme that belongs to the GT 8 family. N-acetylglucosaminyltransferase is a fungal enzyme that catalyzes the addition of N-acetyl-D-glucosamine to mannotetraose side chains by an alpha 1-2 linkage during the synthesis of mannan. The N-acetyl-D-glucosamine moiety in mannan plays a role in the attachment of mannan to asparagine residues in proteins. The mannotetraose and its N-acetyl-D-glucosamine derivative side chains of mannan are the principle immunochemical determinants on the cell surface. N-acetylglucosaminyltransferase is a member of glycosyltransferase family 8, which are, based on the relative anomeric stereochemistry of the substrate and product in the reaction catalyzed, retaining glycosyltransferases. |
| 227884 | Gnt1 | 1.49e-08 | 21 | 158 | 35 | 188 | Alpha-N-acetylglucosamine transferase [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 2 | 513 | 5 | 516 | |
| 3.21e-185 | 42 | 510 | 35 | 515 | |
| 5.99e-56 | 45 | 317 | 64 | 313 | |
| 1.16e-55 | 45 | 319 | 61 | 312 | |
| 1.62e-55 | 39 | 319 | 46 | 302 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.87e-20 | 60 | 214 | 15 | 173 | Crystal structure of human glycogenin-2 catalytic domain [Homo sapiens],4UEG_B Crystal structure of human glycogenin-2 catalytic domain [Homo sapiens] |
|
| 5.84e-18 | 60 | 239 | 16 | 200 | Crystal Structure of Human Glycogenin-1 (GYG1) Tyr195pIPhe mutant, apo form [Homo sapiens],6EQL_A Crystal Structure of Human Glycogenin-1 (GYG1) Tyr195pIPhe mutant complexed with manganese and UDP [Homo sapiens],6EQL_B Crystal Structure of Human Glycogenin-1 (GYG1) Tyr195pIPhe mutant complexed with manganese and UDP [Homo sapiens] |
|
| 6.05e-18 | 36 | 230 | 18 | 212 | Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese [Homo sapiens] |
|
| 1.07e-17 | 60 | 230 | 16 | 191 | Crystal Structure of Human Glycogenin-1 (GYG1), apo form [Homo sapiens],3QVB_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and UDP [Homo sapiens],3U2W_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and glucose or a glucal species [Homo sapiens],3U2W_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and glucose or a glucal species [Homo sapiens] |
|
| 1.07e-17 | 60 | 230 | 16 | 191 | Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and UDP, in a triclinic closed form [Homo sapiens],3T7M_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and UDP, in a triclinic closed form [Homo sapiens],3T7N_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and UDP, in a monoclinic closed form [Homo sapiens],3T7N_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and UDP, in a monoclinic closed form [Homo sapiens],3T7O_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP-Glucose and glucose [Homo sapiens],3T7O_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP-Glucose and glucose [Homo sapiens],3U2U_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and maltotetraose [Homo sapiens],3U2U_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and maltotetraose [Homo sapiens],3U2V_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and maltohexaose [Homo sapiens],3U2V_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and maltohexaose [Homo sapiens],3U2X_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and 1'-deoxyglucose [Homo sapiens],3U2X_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and 1'-deoxyglucose [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.05e-55 | 45 | 325 | 64 | 322 | Putative glucuronosyltransferase PGSIP8 OS=Arabidopsis thaliana OX=3702 GN=PGSIP8 PE=2 SV=1 |
|
| 5.80e-53 | 45 | 403 | 60 | 389 | Putative glucuronosyltransferase PGSIP7 OS=Arabidopsis thaliana OX=3702 GN=PGSIP7 PE=3 SV=1 |
|
| 8.51e-31 | 42 | 320 | 29 | 278 | Inositol phosphorylceramide glucuronosyltransferase 1 OS=Arabidopsis thaliana OX=3702 GN=IPUT1 PE=1 SV=1 |
|
| 5.02e-19 | 39 | 213 | 272 | 447 | Putative UDP-glucuronate:xylan alpha-glucuronosyltransferase 3 OS=Arabidopsis thaliana OX=3702 GN=GUX3 PE=2 SV=1 |
|
| 1.77e-18 | 57 | 250 | 11 | 194 | Glycogenin-1 OS=Caenorhabditis elegans OX=6239 GN=gyg-1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000548 | 0.999421 | CS pos: 20-21. Pr: 0.9724 |
TMHMM Annotations download full data without filtering help
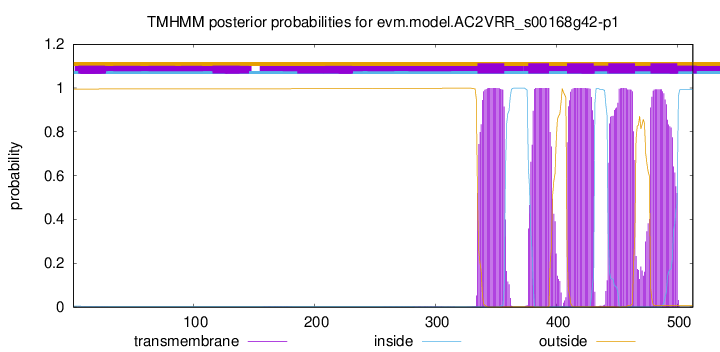
| Start | End |
|---|---|
| 335 | 357 |
| 377 | 394 |
| 409 | 431 |
| 443 | 465 |
| 478 | 500 |
