You are browsing environment: FUNGIDB
CAZyme Information: evm.model.AC2VRR_s00077g19-p1
You are here: Home > Sequence: evm.model.AC2VRR_s00077g19-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Albugo candida | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Albuginaceae; Albugo; Albugo candida | |||||||||||
| CAZyme ID | evm.model.AC2VRR_s00077g19-p1 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 26940; End:33656 Strand: - | |||||||||||
Full Sequence Download help
| MSQPTHYRES SRRRASSNYF ELDADRRRTK SGRKDRLSHR VMSLEAAETI PGDKFSTAPQ | 60 |
| NIVNSTAVLD DDEISIDYCC DFLYAKFGFQ EGSVANQREH VLLLLANGKA RHHPSQPSHH | 120 |
| HITQLHAKLV SNYKSWCEFL KTNPIHYQGV ISGNLRHPLH MEIMLFFLIW GESSNLRHMP | 180 |
| ECICYIFHQL MKHLNQDIQG QDGKKEGWFL ENVVQPIWEE CSNMKRRNHL NKPLEHVKVR | 240 |
| NYDDINEYFW KPYCLKIEVT QVGHELAQKH GKTFYEHRSI FTLILNYYRI FQANILFLTI | 300 |
| LVVLAFAVSI SPNGGRSGFS QFQALGKTIE PFEKRDLKIG FVVLPFVMSL LGICKCVLEF | 360 |
| AHSFHIIFSS ESSLTSSRSW PYTLALAART LWHTGFMALF AFMIYIPLRD QSDTTLLQNA | 420 |
| YAVMAVYIIP GVAILAAQTF YPNLIRKTFA LKFVREGSSS YVGREMAPPW KYKVQYVLFW | 480 |
| IVLWICKSVV SYTILVRPLM LPSLAVYEMR LTYQSSVASF HNILVLVSYW APTVLIFNYD | 540 |
| TQIYFTILQS IIGGYMGWRM KTGEIRGSKE LTRAFRVAPQ LFDQKIVTNL ARSSDLAQSL | 600 |
| NPKDSKTSVN AATYESQMML RFVVVWNEIV NSFREGDLLD DKEAAILQYD IRSNGEVFEP | 660 |
| VFLSAGKLGE AITKTIRNSK DGKSESQLQV SLVEGDCVSA IKSFFTACMY VMEALFGMED | 720 |
| GNVIDGLRMM EEVVENRASM RSFQFQELAR LRLAALDILE EILDLPDPST VSAHLPDTFI | 780 |
| HTLGIIRNFV NKMEVLLNSL QAFSEAPELK GKFVNTKFCS SPNGYMYAAQ GLVNLYRSDV | 840 |
| AMGAATRACL LLSLDRSEAM PRCMEAQRRL GFFMRSLVME IPQLNAIKEM RSFSVVTPFY | 900 |
| AETVLFSLKE LNDPLVNHPI FQKVEEGGKN LTILKYLNKI HPEEWENFLE RVDVASAEEA | 960 |
| QERYPQEIRL WASYRGQTLA RTVQGMMLYE DAIKILHWLE IGSNSARTAE EKQTQLQDMV | 1020 |
| RLKFSYICAC QVYGKHRREN KQQADDIDYL LQEYPNLRVA YVDTIESGEN QYMYDTVLIK | 1080 |
| SEQNEIVEVY RYQLPGDPII GEGKPENQNN AMQFTRGEFV QTIDMNQQHY FEECLKMPQL | 1140 |
| LRTAELHSSG KPVSIIGMRE HIFTGNASSL AKFKTWQELV FVTLSQRVLA DPLYVRMHYG | 1200 |
| HPDIFDKVLA LTRGGVSKSS KGINLSEDVF AGFNATLRGG VVTHVEFMQC GKGRDVALSQ | 1260 |
| ISMFEGKLAN GAGETSLARE AHRMGQFMDF FRLNSMYYSH TGFFFATWMT VVTTFVYMYC | 1320 |
| KVYIALVGVQ DQIIYKMNET SILSQNERYG IPSRAYYDTN AIVNTQYYIQ AGLFLSLPLV | 1380 |
| VVYFSEMGLY RGFFRLIEMV ITGGPFFFIF QVGTTMHYFD NNLVHGEAQY KATGRGFKIT | 1440 |
| RELFVLLYKA YASSHYRKAF ELTGLCLIYL TYGDFNICGP PPTADGNSFS FDFCTTAQSF | 1500 |
| WVQTFAIWFI AITWFIAPYI FNTDGLDFQK TKADIQAWAT WMYADEKYED EDSTMNGGWI | 1560 |
| GWWKSELKLF HNSKPIARLT IVLRESRHFV LMWYVVTLKW DLLTIAYVLG AGVLSVLLLN | 1620 |
| VMCLLRLAFR RCRPIPRALI YVTTVCLAIA AYFAITKYLF STDFQEAASL FYGYIAVLYG | 1680 |
| INEMARMYSF QSTSIANTTI FQELAFFFDF MICFIMIVPL FIMSGIPFLN IVQTRMMYNK | 1740 |
| GFSQIVSASS QYAFSLAAFL GVVGGSGCGW ILYVMSVLES SPGFISYATN YDLLDNKAGD | 1800 |
| GTTTYYFYFA AVMAAVLGGF FNFFFGRRLT VVCGGMLSAL AMVLISAVTT QGESLLFPGT | 1860 |
| ALLGASIGIL LPSIAIYIYE ISTRELRAKS MLLLGTGFII GTMLGSIFMS PEQEIGWIWQ | 1920 |
| LFIANILLSV VTPVVFLFPE SAYWIYMREG ADSCERCLTV LRRKEGVQDE LQLIREESTS | 1980 |
| ANKSTGNIFK FGLGLCLMIV SSLANGALVI YLTTALATSG SSASNFTNCV ILQLVGALLS | 2040 |
| FFVIDRVDLR RVLLGTLLPI TFFVAMLGFN ANTNTWTGDA NNVFLQLVGC LLYFFLGLGT | 2100 |
| STALWASAVG LFPTKQRALY TTFYFMIFFL IPIVALYIRS SPSLYRSAEY VYLYTLAGVC | 2160 |
| ILSTIFLFVA GHLSNGLIGT RQEQEVEEAQ VRRARASRQS SRLTPGGRMR HINRSRDKSY | 2220 |
| NSSGYQLYES PAAGGRLP | 2238 |
Enzyme Prediction help
| EC | 2.4.1.34:28 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT48 | 860 | 1594 | 5.9e-276 | 0.9499323410013532 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396784 | Glucan_synthase | 0.0 | 865 | 1543 | 7 | 701 | 1,3-beta-glucan synthase component. This family consists of various 1,3-beta-glucan synthase components including Gls1, Gls2 and Gls3 from yeast. 1,3-beta-glucan synthase EC:2.4.1.34 also known as callose synthase catalyzes the formation of a beta-1,3-glucan polymer that is a major component of the fungal cell wall. The reaction catalyzed is:- UDP-glucose + {(1,3)-beta-D-glucosyl}(N) <=> UDP + {(1,3)-beta-D-glucosyl}(N+1). |
| 405046 | FKS1_dom1 | 4.07e-29 | 163 | 256 | 6 | 106 | 1,3-beta-glucan synthase subunit FKS1, domain-1. The FKS1_dom1 domain is likely to be the 'Class I' region just N-terminal to the first set of transmembrane helices that is involved in 1,3-beta-glucan synthesis itself. This family is found on proteins with family Glucan_synthase, pfam02364. |
| 340916 | MFS_GLUT6_8_Class3_like | 3.19e-21 | 1756 | 2169 | 2 | 426 | Glucose transporter (GLUT) types 6 and 8, Class 3 GLUTs, and similar transporters of the Major Facilitator Superfamily. This subfamily is composed of glucose transporter type 6 (GLUT6), GLUT8, plant early dehydration-induced gene ERD6-like proteins, and similar insect proteins including facilitated trehalose transporter Tret1-1. GLUTs, also called Solute carrier family 2, facilitated glucose transporters (SLC2A), are a family of proteins that facilitate the transport of hexoses such as glucose and fructose. There are fourteen GLUTs found in humans; they display different substrate specificities and tissue expression. They have been categorized into three classes based on sequence similarity: Class 1 (GLUTs 1-4, 14); Class 2 (GLUTs 5, 7, 9, and 11); and Class 3 (GLUTs 6, 8, 10, 12, and HMIT). Insect Tret1-1 is a low-capacity facilitative transporter for trehalose that mediates the transport of trehalose synthesized in the fat body and the incorporation of trehalose into other tissues that require a carbon source. GLUT proteins are comprised of about 500 amino acid residues, possess a single N-linked oligosaccharide, and have 12 transmembrane segments. They belong to the Glucose transporter -like (GLUT-like) family of the Major Facilitator Superfamily (MFS) of membrane transport proteins. MFS proteins are thought to function through a single substrate binding site, alternating-access mechanism involving a rocker-switch type of movement. |
| 349949 | MFS | 1.63e-13 | 1767 | 2168 | 2 | 378 | Major Facilitator Superfamily. The Major Facilitator Superfamily (MFS) is a large and diverse group of secondary transporters that includes uniporters, symporters, and antiporters. MFS proteins facilitate the transport across cytoplasmic or internal membranes of a variety of substrates including ions, sugar phosphates, drugs, neurotransmitters, nucleosides, amino acids, and peptides. They do so using the electrochemical potential of the transported substrates. Uniporters transport a single substrate, while symporters and antiporters transport two substrates in the same or in opposite directions, respectively, across membranes. MFS proteins are typically 400 to 600 amino acids in length, and the majority contain 12 transmembrane alpha helices (TMs) connected by hydrophilic loops. The N- and C-terminal halves of these proteins display weak similarity and may be the result of a gene duplication/fusion event. Based on kinetic studies and the structures of a few bacterial superfamily members, GlpT (glycerol-3-phosphate transporter), LacY (lactose permease), and EmrD (multidrug transporter), MFS proteins are thought to function through a single substrate binding site, alternating-access mechanism involving a rocker-switch type of movement. Bacterial members function primarily for nutrient uptake, and as drug-efflux pumps to confer antibiotic resistance. Some MFS proteins have medical significance in humans such as the glucose transporter Glut4, which is impaired in type II diabetes, and glucose-6-phosphate transporter (G6PT), which causes glycogen storage disease when mutated. |
| 395036 | Sugar_tr | 1.80e-13 | 1757 | 2169 | 1 | 435 | Sugar (and other) transporter. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CCA17033.1|GT48 | 0.0 | 1 | 2238 | 1 | 2237 |
| CCA22009.1|GT48 | 0.0 | 8 | 2233 | 7 | 2220 |
| UIZ28646.1|GT48 | 0.0 | 12 | 2232 | 14 | 2221 |
| UIZ28580.1|GT48 | 0.0 | 12 | 2232 | 11 | 2236 |
| CCA17037.1|GT48 | 0.0 | 14 | 2232 | 10 | 2241 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| sp|Q9SFU6|CALS9_ARATH | 1.97e-218 | 81 | 1744 | 253 | 1873 | Callose synthase 9 OS=Arabidopsis thaliana OX=3702 GN=CALS9 PE=2 SV=2 |
| sp|Q9SHJ3|CALS7_ARATH | 1.39e-216 | 41 | 1744 | 221 | 1913 | Callose synthase 7 OS=Arabidopsis thaliana OX=3702 GN=CALS7 PE=3 SV=3 |
| sp|Q9SJM0|CALSA_ARATH | 3.31e-216 | 81 | 1744 | 265 | 1887 | Callose synthase 10 OS=Arabidopsis thaliana OX=3702 GN=CALS10 PE=2 SV=5 |
| sp|Q9LXT9|CALS3_ARATH | 3.28e-215 | 81 | 1590 | 239 | 1775 | Callose synthase 3 OS=Arabidopsis thaliana OX=3702 GN=CALS3 PE=3 SV=3 |
| sp|Q9AUE0|CALS1_ARATH | 5.67e-215 | 81 | 1573 | 235 | 1752 | Callose synthase 1 OS=Arabidopsis thaliana OX=3702 GN=CALS1 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000074 | 0.000000 |
TMHMM Annotations download full data without filtering help
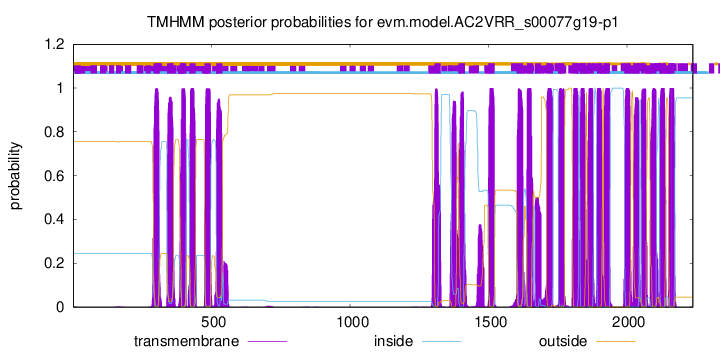
| Start | End |
|---|---|
| 290 | 312 |
| 339 | 361 |
| 386 | 408 |
| 420 | 442 |
| 474 | 496 |
| 517 | 539 |
| 1305 | 1327 |
| 1392 | 1414 |
| 1500 | 1522 |
| 1602 | 1624 |
| 1639 | 1661 |
| 1668 | 1690 |
| 1710 | 1732 |
| 1752 | 1774 |
| 1803 | 1825 |
| 1830 | 1852 |
| 1857 | 1879 |
| 1892 | 1911 |
| 1916 | 1938 |
| 1991 | 2013 |
| 2023 | 2045 |
| 2052 | 2069 |
| 2084 | 2106 |
| 2118 | 2140 |
| 2150 | 2172 |
