You are browsing environment: FUNGIDB
CAZyme Information: Z517_00174-t43_1-p1
You are here: Home > Sequence: Z517_00174-t43_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fonsecaea pedrosoi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Herpotrichiellaceae; Fonsecaea; Fonsecaea pedrosoi | |||||||||||
| CAZyme ID | Z517_00174-t43_1-p1 | |||||||||||
| CAZy Family | AA1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.177:12 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH31 | 182 | 624 | 1e-66 | 0.9976580796252927 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 224418 | YicI | 1.35e-60 | 92 | 684 | 141 | 725 | Alpha-glucosidase, glycosyl hydrolase family GH31 [Carbohydrate transport and metabolism]. |
| 395838 | Glyco_hydro_31 | 1.08e-56 | 183 | 624 | 1 | 442 | Glycosyl hydrolases family 31. Glycosyl hydrolases are key enzymes of carbohydrate metabolism. Family 31 comprises of enzymes that are, or similar to, alpha- galactosidases. |
| 269878 | GH31_NET37 | 1.32e-32 | 270 | 578 | 48 | 361 | glucosidase NET37. NET37 (also known as KIAA1161) is a human lamina-associated nuclear envelope transmembrane protein. A member of the glycosyl hydrolase family 31 (GH31) , it has been shown to be required for myogenic differentiation of C2C12 cells. Related proteins are found in eukaryotes and prokaryotes. Enzymes of the GH31 family possess a wide range of different hydrolytic activities including alpha-glucosidase (glucoamylase and sucrase-isomaltase), alpha-xylosidase, 6-alpha-glucosyltransferase, 3-alpha-isomaltosyltransferase and alpha-1,4-glucan lyase. All GH31 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. |
| 269889 | GH31_GANC_GANAB_alpha | 6.00e-31 | 416 | 630 | 231 | 432 | neutral alpha-glucosidase C, neutral alpha-glucosidase AB. This subgroup includes the closely related glycosyl hydrolase family 31 (GH31) isozymes, neutral alpha-glucosidase C (GANC) and the alpha subunit of heterodimeric neutral alpha-glucosidase AB (GANAB). Initially distinguished on the basis of differences in electrophoretic mobility in starch gel, GANC and GANAB have been shown to have other differences, including those of substrate specificity. GANC and GANAB are key enzymes in glycogen metabolism that hydrolyze terminal, non-reducing 1,4-linked alpha-D-glucose residues from glycogen in the endoplasmic reticulum. The GANC/GANAB family includes the alpha-glucosidase II (ModA) from Dictyostelium discoideum as well as the alpha-glucosidase II (GLS2, or ROT2 - Reversal of TOR2 lethality protein 2) from Saccharomyces cerevisiae. |
| 236731 | PRK10658 | 2.29e-29 | 146 | 621 | 198 | 665 | putative alpha-glucosidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.34e-262 | 26 | 728 | 38 | 725 | |
| 1.48e-44 | 81 | 687 | 134 | 727 | |
| 1.65e-43 | 144 | 627 | 198 | 672 | |
| 1.94e-43 | 78 | 626 | 155 | 695 | |
| 2.16e-43 | 81 | 687 | 134 | 727 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.98e-33 | 55 | 683 | 210 | 865 | Cycloalternan-forming enzyme from Listeria monocytogenes in complex with pentasaccharide substrate [Listeria monocytogenes EGD-e] |
|
| 1.65e-30 | 69 | 631 | 125 | 676 | Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_B Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_C Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_D Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_E Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_F Structure of the YicI thiosugar Michaelis complex [Escherichia coli] |
|
| 1.67e-30 | 69 | 631 | 125 | 676 | Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_B Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_C Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_D Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_E Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_F Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_A Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_B Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_C Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_D Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_E Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_F Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSK_A Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_B Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_C Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_D Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_E Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_F Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli] |
|
| 4.44e-30 | 55 | 683 | 210 | 865 | Cycloalternan-forming enzyme from Listeria monocytogenes in complex with cycloalternan [Listeria monocytogenes EGD-e],5I0D_B Cycloalternan-forming enzyme from Listeria monocytogenes in complex with cycloalternan [Listeria monocytogenes EGD-e] |
|
| 4.52e-30 | 55 | 683 | 231 | 886 | 1.9 Angstrom resolution crystal structure of uncharacterized protein lmo2446 from Listeria monocytogenes EGD-e [Listeria monocytogenes EGD-e],4KWU_A 1.9 Angstrom resolution crystal structure of uncharacterized protein lmo2446 from Listeria monocytogenes EGD-e in complex with alpha-D-glucose, beta-D-glucose, magnesium and calcium [Listeria monocytogenes EGD-e],5HPO_A Cycloalternan-forming enzyme from Listeria monocytogenes in complex with maltopentaose [Listeria monocytogenes EGD-e],5HXM_A Cycloalternan-forming enzyme from Listeria monocytogenes in complex with panose [Listeria monocytogenes] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.87e-34 | 69 | 628 | 123 | 678 | Alpha-xylosidase XylQ OS=Lactiplantibacillus pentosus OX=1589 GN=xylQ PE=1 SV=1 |
|
| 2.98e-32 | 93 | 623 | 167 | 695 | Alpha-xylosidase OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=agdD PE=1 SV=1 |
|
| 8.47e-30 | 69 | 631 | 125 | 676 | Alpha-xylosidase OS=Escherichia coli (strain K12) OX=83333 GN=yicI PE=1 SV=2 |
|
| 4.51e-21 | 134 | 626 | 311 | 812 | Neutral alpha-glucosidase AB OS=Homo sapiens OX=9606 GN=GANAB PE=1 SV=3 |
|
| 2.58e-20 | 387 | 646 | 409 | 656 | Neutral alpha-glucosidase C (Fragment) OS=Macaca fascicularis OX=9541 GN=GANC PE=2 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
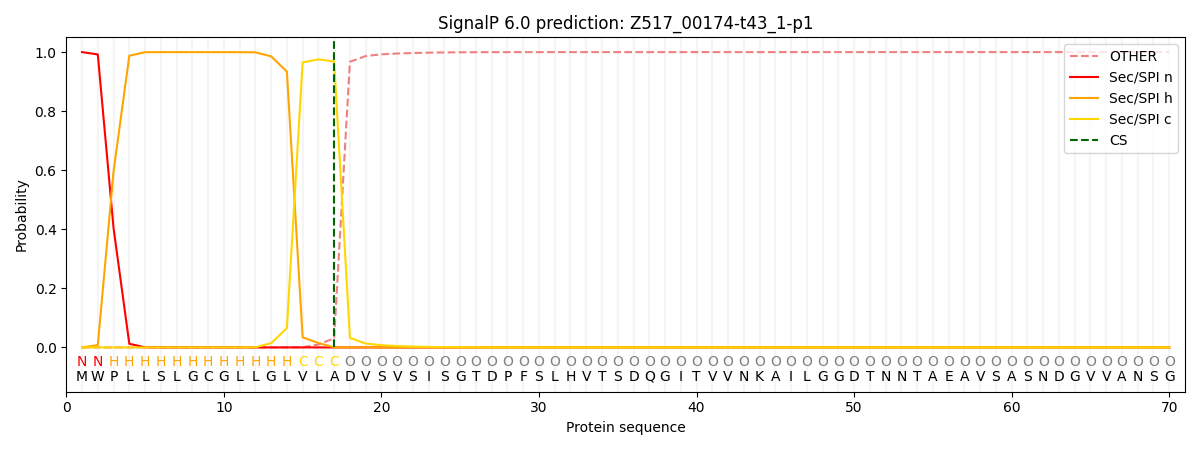
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000244 | 0.999714 | CS pos: 17-18. Pr: 0.9685 |
