You are browsing environment: FUNGIDB
CAZyme Information: XP_001912538.1
You are here: Home > Sequence: XP_001912538.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
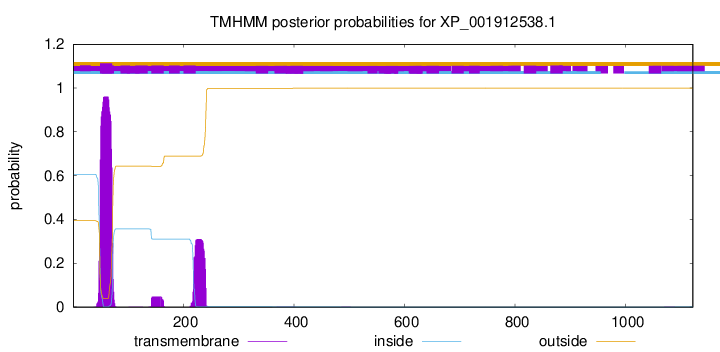
TMHMM annotations
Basic Information help
| Species | Podospora anserina | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Podosporaceae; Podospora; Podospora anserina | |||||||||||
| CAZyme ID | XP_001912538.1 | |||||||||||
| CAZy Family | GT22 | |||||||||||
| CAZyme Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase [Source:UniProtKB/TrEMBL;Acc:B2AAA8] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 89 | 643 | 9.9e-138 | 0.9955156950672646 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 7.42e-130 | 89 | 644 | 1 | 453 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 7.98e-57 | 71 | 648 | 61 | 522 | glycoside hydrolase family 47 protein; Provisional |
| 239039 | PA_PoS1_like | 2.98e-05 | 961 | 1018 | 11 | 78 | PA_PoS1_like: Protease-associated (PA) domain PoS1-like. This group includes various PA domain-containing proteins similar to Pleurotus ostreatus (Po)S1. PoSl, the main extracellular protease in P. ostreatus is a subtilisin-like serine protease belonging to the peptidase S8 family. Ca2+ and Mn2+ both stimulate the protease activity of (Po)S1. Ca2+ protects PoS1 from autolysis. PoS1 is a monomeric glycoprotein, which may play a role in the regulation of laccases in lignin formation. (Po)S1 participates in the degradation of POXA1b, and in the activation of POXA3, (POXA1b and POXA3 are laccase isoenzymes), but its effect may be indirect. The significance of the PA domain to PoS1 has not been ascertained. It may be a protein-protein interaction domain. At peptidase active sites, the PA domain may participate in substrate binding and/or promoting conformational changes, which influence the stability and accessibility of the site to substrate. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 1122 | 1 | 1122 | |
| 0.0 | 1 | 1122 | 1 | 1157 | |
| 0.0 | 1 | 1122 | 1 | 1144 | |
| 0.0 | 45 | 1122 | 35 | 1112 | |
| 0.0 | 74 | 1117 | 1 | 1023 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.82e-32 | 87 | 644 | 10 | 451 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase and Man9GlcNAc2-PA complex [Homo sapiens] |
|
| 2.00e-32 | 87 | 644 | 15 | 456 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase [Homo sapiens] |
|
| 2.06e-32 | 87 | 644 | 15 | 456 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With 1-Deoxymannojirimycin [Homo sapiens],1FO3_A Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With Kifunensine [Homo sapiens] |
|
| 5.78e-32 | 87 | 644 | 93 | 534 | Crystal Structure Of Human Class I alpha-1,2-Mannosidase In Complex With Thio-Disaccharide Substrate Analogue [Homo sapiens] |
|
| 8.39e-32 | 87 | 644 | 10 | 451 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens],5KK7_B Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.67e-118 | 58 | 716 | 11 | 539 | ER degradation-enhancing alpha-mannosidase-like protein 1 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=mnl1 PE=3 SV=2 |
|
| 5.05e-114 | 81 | 646 | 124 | 583 | ER degradation-enhancing alpha-mannosidase-like protein 1 OS=Mus musculus OX=10090 GN=Edem1 PE=1 SV=1 |
|
| 1.60e-112 | 81 | 644 | 129 | 586 | ER degradation-enhancing alpha-mannosidase-like protein 1 OS=Homo sapiens OX=9606 GN=EDEM1 PE=1 SV=1 |
|
| 6.99e-100 | 56 | 646 | 5 | 504 | ER degradation-enhancing alpha-mannosidase-like protein 1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=MNL1 PE=1 SV=1 |
|
| 2.90e-91 | 80 | 644 | 35 | 478 | Alpha-mannosidase I MNS5 OS=Arabidopsis thaliana OX=3702 GN=MNS5 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999966 | 0.000054 |