You are browsing environment: FUNGIDB
CAZyme Information: XP_001910674.1
You are here: Home > Sequence: XP_001910674.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Podospora anserina | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Podosporaceae; Podospora; Podospora anserina | |||||||||||
| CAZyme ID | XP_001910674.1 | |||||||||||
| CAZy Family | GH67 | |||||||||||
| CAZyme Description | GMC_OxRdtase_N domain-containing protein [Source:UniProtKB/TrEMBL;Acc:B2B3X4] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA3 | 38 | 650 | 4.5e-148 | 0.9947183098591549 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 225186 | BetA | 7.02e-67 | 37 | 651 | 6 | 536 | Choline dehydrogenase or related flavoprotein [Lipid transport and metabolism, General function prediction only]. |
| 235000 | PRK02106 | 2.75e-59 | 37 | 648 | 4 | 531 | choline dehydrogenase; Validated |
| 398739 | GMC_oxred_C | 8.93e-39 | 506 | 644 | 1 | 143 | GMC oxidoreductase. This domain found associated with pfam00732. |
| 369159 | HET | 2.86e-32 | 889 | 1036 | 1 | 146 | Heterokaryon incompatibility protein (HET). This family represents a conserved region approximately 150 residues long within various heterokaryon incompatibility proteins that seem to be restricted to ascomycete fungi. Genetic differences in specific het genes prevent a viable heterokaryotic fungal cell from being formed by the fusion of filaments from two different wild-type strains. Many family members also contain the pfam00400 repeat and the pfam05729 domain. |
| 366272 | GMC_oxred_N | 2.54e-22 | 129 | 372 | 15 | 217 | GMC oxidoreductase. This family of proteins bind FAD as a cofactor. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 1389 | 1 | 1370 | |
| 0.0 | 1 | 653 | 1 | 634 | |
| 8.46e-248 | 10 | 655 | 7 | 643 | |
| 5.75e-197 | 152 | 652 | 3 | 507 | |
| 3.85e-115 | 29 | 648 | 34 | 620 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.07e-253 | 11 | 652 | 5 | 642 | Structural insights on a new fungal aryl-alcohol oxidase [Thermothelomyces thermophilus],6O9N_B Structural insights on a new fungal aryl-alcohol oxidase [Thermothelomyces thermophilus] |
|
| 2.32e-90 | 35 | 653 | 2 | 587 | Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE2_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE3_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE4_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE4_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE5_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE5_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE6_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE6_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE7_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE7_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],7AA2_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],7AA2_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495] |
|
| 3.96e-56 | 37 | 648 | 15 | 587 | Chain A, Oligosaccharide dehydrogenase [Trametes cinnabarina],6XUU_A Chain A, Oligosaccharide dehydrogenase [Trametes cinnabarina],6XUV_A Chain A, Oligosaccharide dehydrogenase [Trametes cinnabarina] |
|
| 1.65e-48 | 38 | 648 | 5 | 564 | Crystal structure of Aspergillus flavus FAD glucose dehydrogenase [Aspergillus flavus NRRL3357],4YNU_A Crystal structure of Aspergillus flavus FADGDH in complex with D-glucono-1,5-lactone [Aspergillus flavus NRRL3357] |
|
| 1.29e-41 | 40 | 652 | 21 | 579 | GLUCOSE OXIDASE FROM APERGILLUS NIGER [Aspergillus niger],1GAL_A CRYSTAL STRUCTURE OF GLUCOSE OXIDASE FROM ASPERGILLUS NIGER: REFINED AT 2.3 ANGSTROMS RESOLUTION [Aspergillus niger],3QVP_A Crystal structure of glucose oxidase for space group C2221 at 1.2 A resolution [Aspergillus niger],3QVR_A Crystal structure of glucose oxidase for space group P3121 at 1.3 A resolution. [Aspergillus niger] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.03e-83 | 25 | 648 | 24 | 613 | Dehydrogenase xptC OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=xptC PE=3 SV=1 |
|
| 1.38e-67 | 37 | 649 | 45 | 616 | GMC oxidoreductase family protein Mala s 12.0101 OS=Malassezia sympodialis OX=76777 PE=1 SV=2 |
|
| 1.38e-67 | 37 | 649 | 45 | 616 | GMC oxidoreductase family protein Mala s 12 OS=Malassezia sympodialis (strain ATCC 42132) OX=1230383 GN=MSY001_2108 PE=1 SV=1 |
|
| 2.96e-55 | 39 | 651 | 36 | 610 | Oxidoreductase cicC OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=cicC PE=1 SV=1 |
|
| 1.76e-52 | 38 | 651 | 39 | 632 | Dehydrogenase ARMGADRAFT_1018426 OS=Armillaria gallica OX=47427 GN=ARMGADRAFT_1018426 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
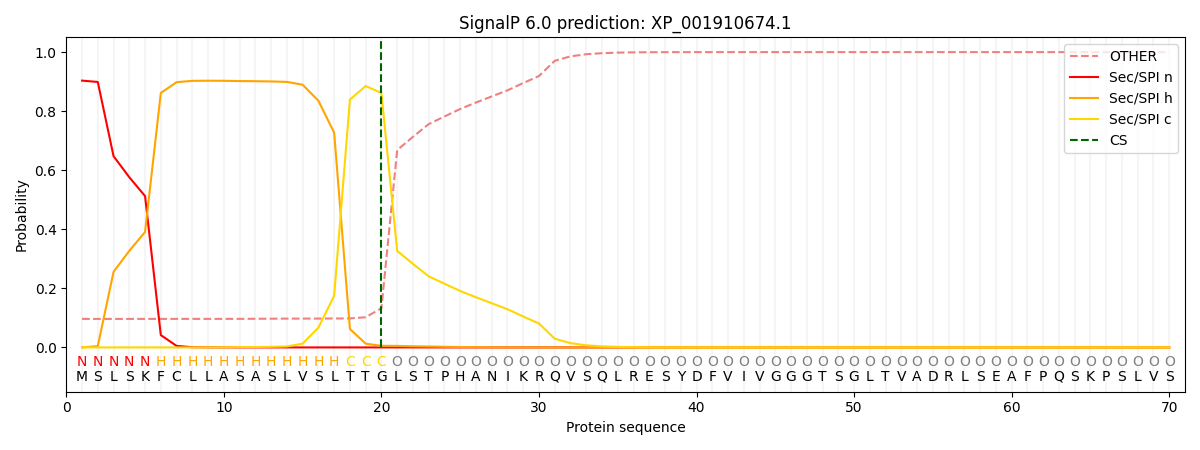
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.100846 | 0.899143 | CS pos: 20-21. Pr: 0.8617 |
