You are browsing environment: FUNGIDB
CAZyme Information: XP_001907222.1
You are here: Home > Sequence: XP_001907222.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Podospora anserina | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Podosporaceae; Podospora; Podospora anserina | |||||||||||
| CAZyme ID | XP_001907222.1 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | Podospora anserina S mat+ genomic DNA chromosome 1, supercontig 4 [Source:UniProtKB/TrEMBL;Acc:B2AU20] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH55 | 89 | 820 | 7.4e-257 | 0.9594594594594594 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 403800 | Pectate_lyase_3 | 7.64e-72 | 117 | 353 | 1 | 213 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| 173787 | Peptidases_S8_S53 | 8.89e-18 | 1500 | 1761 | 2 | 234 | Peptidase domain in the S8 and S53 families. Members of the peptidases S8 (subtilisin and kexin) and S53 (sedolisin) family include endopeptidases and exopeptidases. The S8 family has an Asp/His/Ser catalytic triad similar to that found in trypsin-like proteases, but do not share their three-dimensional structure and are not homologous to trypsin. Serine acts as a nucleophile, aspartate as an electrophile, and histidine as a base. The S53 family contains a catalytic triad Glu/Asp/Ser with an additional acidic residue Asp in the oxyanion hole, similar to that of subtilisin. The serine residue here is the nucleophilic equivalent of the serine residue in the S8 family, while glutamic acid has the same role here as the histidine base. However, the aspartic acid residue that acts as an electrophile is quite different. In S53, it follows glutamic acid, while in S8 it precedes histidine. The stability of these enzymes may be enhanced by calcium; some members have been shown to bind up to 4 ions via binding sites with different affinity. There is a great diversity in the characteristics of their members: some contain disulfide bonds, some are intracellular while others are extracellular, some function at extreme temperatures, and others at high or low pH values. |
| 173790 | Peptidases_S8_PCSK9_ProteinaseK_like | 9.42e-17 | 1479 | 1772 | 10 | 250 | Peptidase S8 family domain in ProteinaseK-like proteins. The peptidase S8 or Subtilase clan of proteases have a Asp/His/Ser catalytic triad that is not homologous to trypsin. This CD contains several members of this clan including: PCSK9 (Proprotein convertase subtilisin/kexin type 9), Proteinase_K, Proteinase_T, and other subtilisin-like serine proteases. PCSK9 posttranslationally regulates hepatic low-density lipoprotein receptors (LDLRs) by binding to LDLRs on the cell surface, leading to their degradation. The binding site of PCSK9 has been localized to the epidermal growth factor-like repeat A (EGF-A) domain of the LDLR. Characterized Proteinases K are secreted endopeptidases with a high degree of sequence conservation. Proteinases K are not substrate-specific and function in a wide variety of species in different pathways. It can hydrolyze keratin and other proteins with subtilisin-like specificity. The number of calcium-binding motifs found in these differ. Proteinase T is a novel proteinase from the fungus Tritirachium album Limber. The amino acid sequence of proteinase T as deduced from the nucleotide sequence is about 56% identical to that of proteinase K. |
| 173794 | Peptidases_S8_Autotransporter_serine_protease_like | 5.83e-13 | 1511 | 1748 | 16 | 244 | Peptidase S8 family domain in Autotransporter serine proteases. Autotransporter serine proteases belong to Peptidase S8 or Subtilase family. Subtilases, or subtilisin-like serine proteases, have an Asp/His/Ser catalytic triad similar to that found in trypsin-like proteases, but do not share their three-dimensional structure (an example of convergent evolution). Autotransporters are a superfamily of outer membrane/secreted proteins of gram-negative bacteria. The presence of these subtilisin-like domains in these autotransporters are may enable them to be auto-catalytic and may also serve to allow them to act as a maturation protease cleaving other outer membrane proteins at the cell surface. |
| 173809 | Peptidases_S8_Subtilisin_Novo-like | 4.58e-10 | 1612 | 1752 | 144 | 272 | Peptidase S8 family domain in Subtilisin_Novo-like proteins. Subtilisins are a group of alkaline proteinases originating from different strains of Bacillus subtilis. Novo is one of the strains that produced enzymes belonging to this group. The enzymes obtained from the Novo and BPN' strains are identical. The Carlsburg and Novo subtilisins are thought to have arisen from a common ancestral protein. They have similar peptidase and esterase activities, pH profiles, catalyze transesterification reactions, and are both inhibited by diispropyl fluorophosphate, though they differ in 85 positions in the amino acid sequence. Members of the peptidases S8 and S35 clan include endopeptidases, exopeptidases and also a tripeptidyl-peptidase. The S8 family has an Asp/His/Ser catalytic triad similar to that found in trypsin-like proteases, but do not share their three-dimensional structure and are not homologous to trypsin. The S53 family contains a catalytic triad Glu/Asp/Ser with an additional acidic residue Asp in the oxyanion hole, similar to that of subtilisin.. The stability of these enzymes may be enhanced by calcium, some members have been shown to bind up to 4 ions via binding sites with different affinity. Some members of this clan contain disulfide bonds. These enzymes can be intra- and extracellular, some function at extreme temperatures and pH values. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 2168 | 1 | 2168 | |
| 0.0 | 1 | 2168 | 1 | 2172 | |
| 0.0 | 84 | 2018 | 24 | 1906 | |
| 0.0 | 1 | 1617 | 1 | 1548 | |
| 0.0 | 1 | 1207 | 42 | 1190 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.17e-179 | 93 | 816 | 4 | 728 | Chain A, Beta-1,3-glucanase [Thermochaetoides thermophila],5M60_A Chain A, Beta-1,3-glucanase [Thermochaetoides thermophila] |
|
| 1.57e-169 | 81 | 813 | 15 | 727 | Chain A, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQN_B Chain B, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQO_A Chain A, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQO_B Chain B, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium] |
|
| 1.59e-06 | 1533 | 1753 | 59 | 244 | Crystal structure of Penicillium cyclopium protease [Penicillium cyclopium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.38e-168 | 91 | 816 | 39 | 848 | Probable glucan endo-1,3-beta-glucosidase ARB_02077 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_02077 PE=1 SV=1 |
|
| 2.12e-150 | 95 | 812 | 48 | 755 | Glucan 1,3-beta-glucosidase OS=Cochliobolus carbonum OX=5017 GN=EXG1 PE=1 SV=1 |
|
| 1.14e-53 | 90 | 815 | 32 | 730 | Glucan endo-1,3-beta-glucosidase BGN13.1 OS=Trichoderma harzianum OX=5544 GN=bgn13.1 PE=1 SV=1 |
|
| 1.04e-08 | 1473 | 1771 | 125 | 388 | Subtilisin-like protease 3 OS=Pseudogymnoascus destructans (strain ATCC MYA-4855 / 20631-21) OX=658429 GN=SP3 PE=1 SV=1 |
|
| 1.35e-08 | 1483 | 1771 | 141 | 374 | Subtilisin-like protease 2 OS=Pseudogymnoascus destructans (strain ATCC MYA-4855 / 20631-21) OX=658429 GN=SP2 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
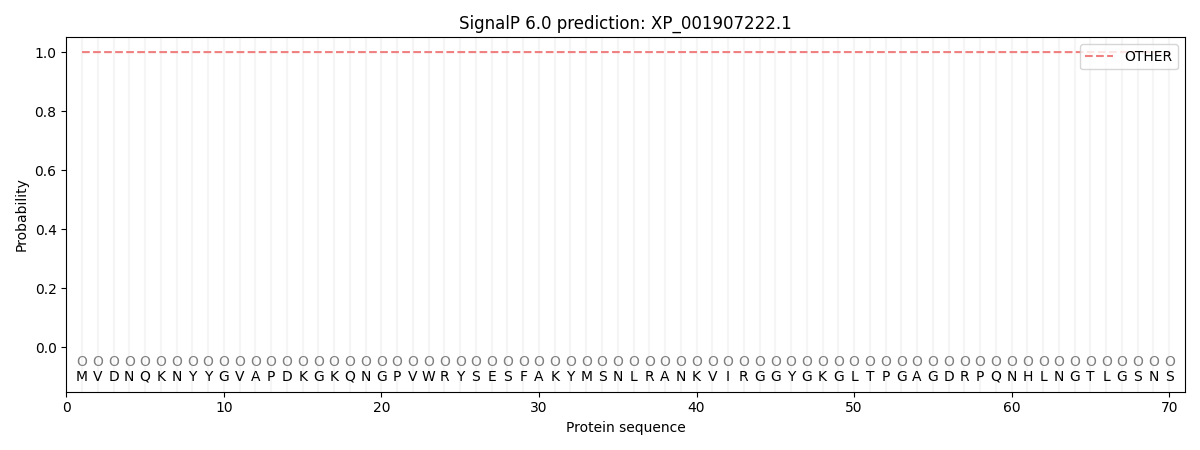
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000041 | 0.000001 |
