You are browsing environment: FUNGIDB
CAZyme Information: XP_001905511.1
You are here: Home > Sequence: XP_001905511.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Podospora anserina | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Podosporaceae; Podospora; Podospora anserina | |||||||||||
| CAZyme ID | XP_001905511.1 | |||||||||||
| CAZy Family | CBM52 | |||||||||||
| CAZyme Description | Podospora anserina S mat+ genomic DNA chromosome 7, supercontig 3 [Source:UniProtKB/TrEMBL;Acc:B2AP59] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT55 | 38 | 435 | 2.5e-136 | 0.9738903394255874 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 401440 | Osmo_MPGsynth | 4.90e-147 | 65 | 435 | 31 | 380 | Mannosyl-3-phosphoglycerate synthase (osmo_MPGsynth). This family consists of examples of mannosyl-3-phosphoglycerate synthase (MPGS), which together with mannosyl-3-phosphoglycerate phosphatase (MPGP) EC:2.4.1.217, comprises a two-step pathway for mannosylglycerate biosynthesis. Mannosylglycerate is a compatible solute that tends to be restricted to extreme thermophiles of archaea and bacteria. Note that in Rhodothermus marinus, this pathway is one of two; the other is condensation of GDP-mannose with D-glycerate by mannosylglycerate synthase. |
| 237735 | PRK14503 | 4.96e-93 | 76 | 437 | 44 | 383 | mannosyl-3-phosphoglycerate synthase; Provisional |
| 162866 | osmo_MPGsynth | 6.83e-83 | 63 | 435 | 30 | 380 | mannosyl-3-phosphoglycerate synthase. This family consists of examples of mannosyl-3-phosphoglycerate synthase (MPGS), which together mannosyl-3-phosphoglycerate phosphatase (MPGP) comprises a two-step pathway for mannosylglycerate biosynthesis. Mannosylglycerate is a compatible solute that tends to be restricted to extreme thermophiles of archaea and bacteria. Note that in Rhodothermus marinus, this pathway is one of two; the other is condensation of GDP-mannose with D-glycerate by mannosylglycerate synthase. |
| 184713 | PRK14502 | 2.69e-71 | 78 | 435 | 50 | 385 | bifunctional mannosyl-3-phosphoglycerate synthase/mannosyl-3 phosphoglycerate phosphatase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 502 | 1 | 502 | |
| 0.0 | 1 | 502 | 1 | 502 | |
| 0.0 | 1 | 502 | 1 | 502 | |
| 8.73e-88 | 1 | 435 | 81 | 611 | |
| 6.41e-85 | 1 | 456 | 65 | 549 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.74e-52 | 81 | 435 | 49 | 382 | Crystal structure of mannosyl-3-phosphoglycerate synthase from Pyrococcus horikoshii [Pyrococcus horikoshii],2ZU7_B Crystal structure of mannosyl-3-phosphoglycerate synthase from Pyrococcus horikoshii [Pyrococcus horikoshii],2ZU8_A Crystal structure of mannosyl-3-phosphoglycerate synthase from Pyrococcus horikoshii [Pyrococcus horikoshii],2ZU8_B Crystal structure of mannosyl-3-phosphoglycerate synthase from Pyrococcus horikoshii [Pyrococcus horikoshii],2ZU9_A Crystal structure of mannosyl-3-phosphoglycerate synthase from Pyrococcus horikoshii [Pyrococcus horikoshii],2ZU9_B Crystal structure of mannosyl-3-phosphoglycerate synthase from Pyrococcus horikoshii [Pyrococcus horikoshii] |
|
| 3.55e-52 | 77 | 435 | 47 | 381 | Mannosyl-3-phosphoglycerate synthase from Thermus thermophilus HB27 apoprotein [Thermus thermophilus HB27],2WVK_B Mannosyl-3-phosphoglycerate synthase from Thermus thermophilus HB27 apoprotein [Thermus thermophilus HB27],2WVL_A Mannosyl-3-phosphoglycerate synthase from Thermus thermophilus HB27 in complex with GDP-alpha-D-Mannose and Mg(II) [Thermus thermophilus HB27],2WVL_B Mannosyl-3-phosphoglycerate synthase from Thermus thermophilus HB27 in complex with GDP-alpha-D-Mannose and Mg(II) [Thermus thermophilus HB27] |
|
| 9.89e-51 | 77 | 435 | 47 | 381 | H309A mutant of Mannosyl-3-phosphoglycerate synthase from Thermus thermophilus HB27 in complex with GDP-alpha-D-Mannose and Mg(II) [Thermus thermophilus HB27],2WVM_B H309A mutant of Mannosyl-3-phosphoglycerate synthase from Thermus thermophilus HB27 in complex with GDP-alpha-D-Mannose and Mg(II) [Thermus thermophilus HB27] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.96e-58 | 70 | 437 | 37 | 384 | Mannosyl-3-phosphoglycerate synthase OS=Pyrococcus abyssi (strain GE5 / Orsay) OX=272844 GN=mngA PE=3 SV=1 |
|
| 4.27e-57 | 81 | 435 | 49 | 382 | Mannosyl-3-phosphoglycerate synthase OS=Pyrococcus horikoshii (strain ATCC 700860 / DSM 12428 / JCM 9974 / NBRC 100139 / OT-3) OX=70601 GN=mngA PE=1 SV=1 |
|
| 6.76e-54 | 82 | 435 | 50 | 382 | Mannosyl-3-phosphoglycerate synthase OS=Pyrococcus furiosus (strain ATCC 43587 / DSM 3638 / JCM 8422 / Vc1) OX=186497 GN=mngA PE=3 SV=1 |
|
| 6.19e-38 | 77 | 444 | 44 | 386 | Mannosyl-3-phosphoglycerate synthase OS=Aeropyrum pernix (strain ATCC 700893 / DSM 11879 / JCM 9820 / NBRC 100138 / K1) OX=272557 GN=mngA PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
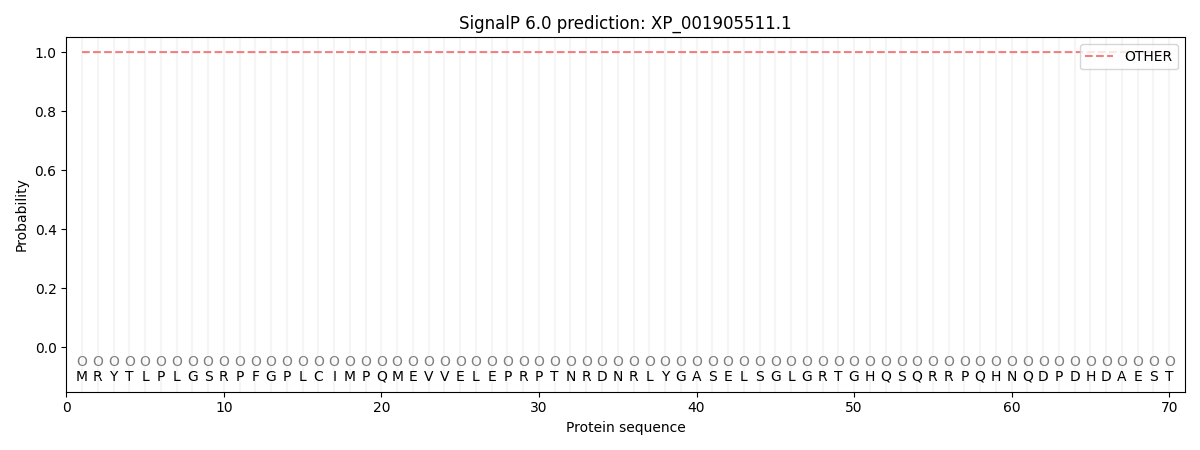
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000044 | 0.000000 |
