You are browsing environment: FUNGIDB
CAZyme Information: XP_001903674.1
You are here: Home > Sequence: XP_001903674.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Podospora anserina | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Podosporaceae; Podospora; Podospora anserina | |||||||||||
| CAZyme ID | XP_001903674.1 | |||||||||||
| CAZy Family | AA3 | |||||||||||
| CAZyme Description | Podospora anserina S mat+ genomic DNA chromosome 4, supercontig 1 [Source:UniProtKB/TrEMBL;Acc:B2ADD4] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH125 | 1 | 387 | 1.9e-140 | 0.8980099502487562 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 399660 | Glyco_hydro_125 | 0.0 | 1 | 386 | 42 | 415 | Metal-independent alpha-mannosidase (GH125). This family, which contains bacterial and fungal glycoside hydrolases, is also known as GH125. They function as metal-independent alpha-mannosidases, with specificity for alpha-1,6-linked non-reducing terminal mannose residues. Structurally this family is part of the 6 hairpin glycosidase superfamily. |
| 226068 | COG3538 | 9.36e-127 | 1 | 393 | 61 | 426 | Meiotically up-regulated gene 157 (Mug157) protein (function unknown) [Function unknown]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.69e-296 | 1 | 402 | 1 | 402 | |
| 3.69e-296 | 1 | 402 | 1 | 402 | |
| 2.22e-288 | 1 | 402 | 123 | 524 | |
| 9.84e-152 | 1 | 402 | 120 | 528 | |
| 1.73e-150 | 1 | 402 | 157 | 563 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.42e-73 | 2 | 402 | 107 | 481 | Crystal structure of BT3781 protein from Bacteroides thetaiotaomicron, Northeast Structural Genomics Target BtR58 [Bacteroides thetaiotaomicron VPI-5482],2P0V_B Crystal structure of BT3781 protein from Bacteroides thetaiotaomicron, Northeast Structural Genomics Target BtR58 [Bacteroides thetaiotaomicron VPI-5482] |
|
| 1.31e-71 | 2 | 402 | 87 | 461 | Crystal structure of an exo-alpha-1,6-mannosidase (bacova_03626) from bacteroides ovatus at 1.70 a resolution [Bacteroides ovatus ATCC 8483],3ON6_B Crystal structure of an exo-alpha-1,6-mannosidase (bacova_03626) from bacteroides ovatus at 1.70 a resolution [Bacteroides ovatus ATCC 8483] |
|
| 1.93e-71 | 2 | 402 | 89 | 463 | Crystal structure of an exo-alpha-1,6-mannosidase (bacova_03347) from bacteroides ovatus at 1.60 a resolution [Bacteroides ovatus ATCC 8483],3P2C_B Crystal structure of an exo-alpha-1,6-mannosidase (bacova_03347) from bacteroides ovatus at 1.60 a resolution [Bacteroides ovatus ATCC 8483] |
|
| 1.86e-67 | 1 | 394 | 61 | 427 | Crystal structure of GH125 1,6-alpha-mannosidase from Clostridium perfringens in complex with mannoimidazole [Clostridium perfringens str. 13],6RQK_B Crystal structure of GH125 1,6-alpha-mannosidase from Clostridium perfringens in complex with mannoimidazole [Clostridium perfringens str. 13] |
|
| 2.99e-67 | 1 | 394 | 61 | 427 | Analysis of a New Family of Widely Distributed Metal-independent alpha-Mannosidases Provides Unique Insight into the Processing of N-linked Glycans, Clostridium perfringens CPE0426 apo-structure [Clostridium perfringens],3QT9_A Analysis of a new family of widely distributed metal-independent alpha mannosidases provides unique insight into the processing of N-linked glycans, Clostridium perfringens CPE0426 complexed with alpha-1,6-linked 1-thio-alpha-mannobiose [Clostridium perfringens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.97e-105 | 2 | 400 | 119 | 507 | Meiotically up-regulated gene 157 protein OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=mug157 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
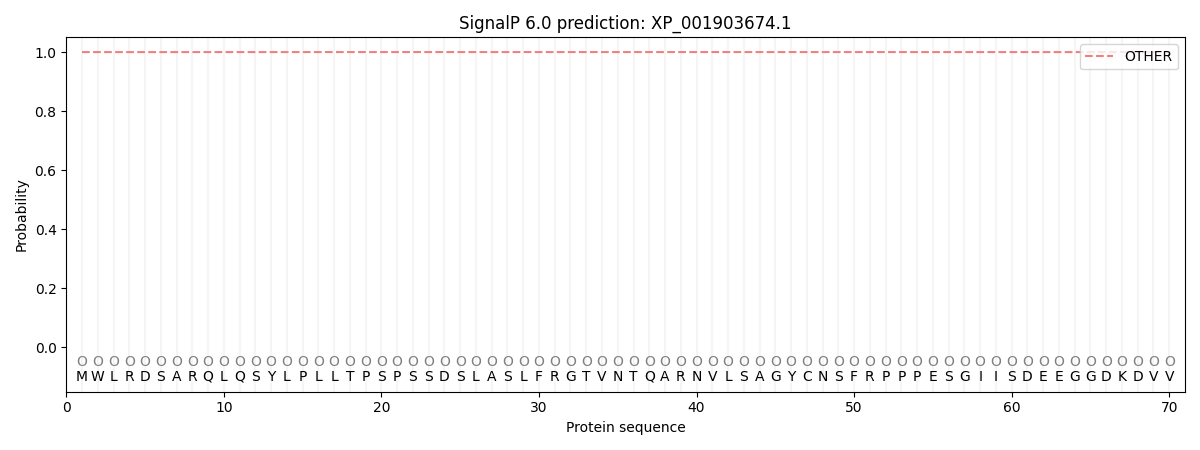
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000079 | 0.000000 |
