You are browsing environment: FUNGIDB
CAZyme Information: VDB93860.1
You are here: Home > Sequence: VDB93860.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Blumeria graminis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Leotiomycetes; ; Erysiphaceae; Blumeria; Blumeria graminis | |||||||||||
| CAZyme ID | VDB93860.1 | |||||||||||
| CAZy Family | GT39 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH128 | 213 | 421 | 1.2e-65 | 0.9776785714285714 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 371727 | Glyco_hydro_cc | 3.10e-75 | 203 | 422 | 1 | 235 | Glycosyl hydrolase catalytic core. This family is probably a glycosyl hydrolase, and is conserved in fungi and some Proteobacteria. The pombe member is annotated as being from IPR013781. |
| 223021 | PHA03247 | 0.001 | 51 | 195 | 2722 | 2888 | large tegument protein UL36; Provisional |
| 237863 | PRK14949 | 0.001 | 77 | 191 | 651 | 765 | DNA polymerase III subunits gamma and tau; Provisional |
| 237057 | PRK12323 | 0.002 | 78 | 190 | 449 | 567 | DNA polymerase III subunit gamma/tau. |
| 235124 | PRK03427 | 0.005 | 125 | 193 | 108 | 174 | cell division protein ZipA; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 9.96e-271 | 1 | 424 | 1 | 424 | |
| 9.45e-85 | 197 | 422 | 178 | 408 | |
| 9.68e-81 | 198 | 422 | 177 | 406 | |
| 9.68e-81 | 198 | 422 | 177 | 406 | |
| 1.88e-78 | 199 | 422 | 182 | 410 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.59e-63 | 197 | 422 | 2 | 227 | Crystal structure of a GH128 (subgroup III) curdlan-specific exo-beta-1,3-glucanase from Blastomyces gilchristii (BgGH128_III) in complex with laminaribiose at -2 and -1 subsites [Blastomyces gilchristii SLH14081] |
|
| 2.50e-62 | 197 | 422 | 33 | 258 | Crystal structure of a GH128 (subgroup III) curdlan-specific exo-beta-1,3-glucanase from Blastomyces gilchristii (BgGH128_III) [Blastomyces gilchristii],6UAY_B Crystal structure of a GH128 (subgroup III) curdlan-specific exo-beta-1,3-glucanase from Blastomyces gilchristii (BgGH128_III) [Blastomyces gilchristii],6UAZ_A Crystal structure of a GH128 (subgroup III) curdlan-specific exo-beta-1,3-glucanase from Blastomyces gilchristii (BgGH128_III) in complex with glucose [Blastomyces gilchristii],6UAZ_B Crystal structure of a GH128 (subgroup III) curdlan-specific exo-beta-1,3-glucanase from Blastomyces gilchristii (BgGH128_III) in complex with glucose [Blastomyces gilchristii],6UB1_A Crystal structure of a GH128 (subgroup III) curdlan-specific exo-beta-1,3-glucanase from Blastomyces gilchristii (BgGH128_III) in complex with laminaribiose at -3 and -2 subsites [Blastomyces gilchristii],6UB1_B Crystal structure of a GH128 (subgroup III) curdlan-specific exo-beta-1,3-glucanase from Blastomyces gilchristii (BgGH128_III) in complex with laminaribiose at -3 and -2 subsites [Blastomyces gilchristii],6UB1_C Crystal structure of a GH128 (subgroup III) curdlan-specific exo-beta-1,3-glucanase from Blastomyces gilchristii (BgGH128_III) in complex with laminaribiose at -3 and -2 subsites [Blastomyces gilchristii],6UB1_D Crystal structure of a GH128 (subgroup III) curdlan-specific exo-beta-1,3-glucanase from Blastomyces gilchristii (BgGH128_III) in complex with laminaribiose at -3 and -2 subsites [Blastomyces gilchristii] |
|
| 3.38e-26 | 199 | 424 | 95 | 344 | Crystal structure of a GH128 (subgroup II) endo-beta-1,3-glucanase from Sorangium cellulosum (ScGH128_II) [Sorangium cellulosum So ce56],6UAX_B Crystal structure of a GH128 (subgroup II) endo-beta-1,3-glucanase from Sorangium cellulosum (ScGH128_II) [Sorangium cellulosum So ce56] |
|
| 1.86e-23 | 190 | 422 | 19 | 259 | Crystal structure of a GH128 (subgroup I) endo-beta-1,3-glucanase from Amycolatopsis mediterranei (AmGH128_I) [Amycolatopsis mediterranei],6UAR_A Crystal structure of a GH128 (subgroup I) endo-beta-1,3-glucanase from Amycolatopsis mediterranei (AmGH128_I) in complex with laminaritriose [Amycolatopsis mediterranei] |
|
| 4.81e-23 | 190 | 422 | 19 | 259 | Crystal structure of a GH128 (subgroup I) endo-beta-1,3-glucanase (E199Q mutant) from Amycolatopsis mediterranei (AmGH128_I) in the complex with laminarihexaose [Amycolatopsis mediterranei],6UFZ_A Crystal structure of a GH128 (subgroup I) endo-beta-1,3-glucanase (E199Q mutant) from Amycolatopsis mediterranei (AmGH128_I) [Amycolatopsis mediterranei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.49e-26 | 194 | 424 | 291 | 530 | Alkali-sensitive linkage protein 1 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=asl1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
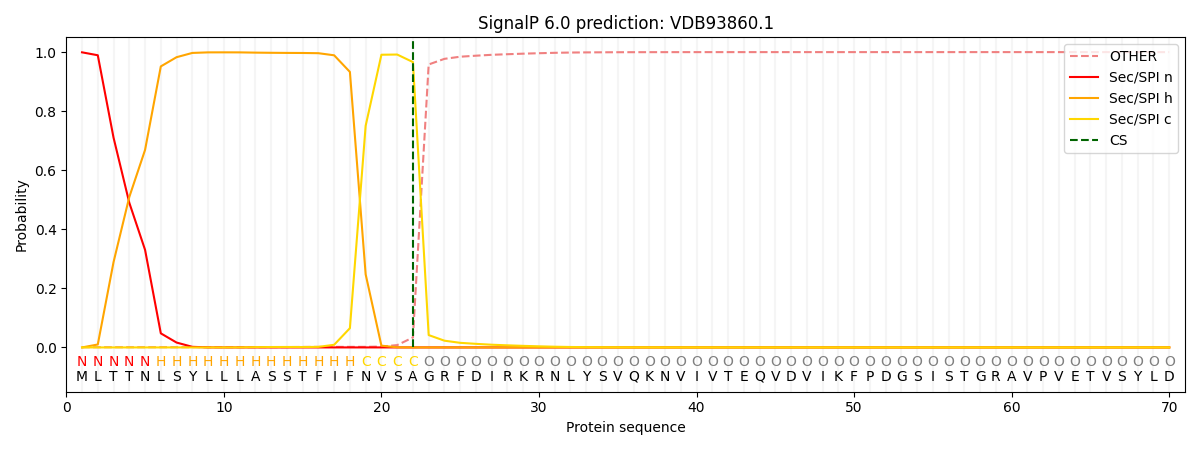
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.001710 | 0.998261 | CS pos: 22-23. Pr: 0.9663 |
