You are browsing environment: FUNGIDB
CAZyme Information: VDB86103.1
You are here: Home > Sequence: VDB86103.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
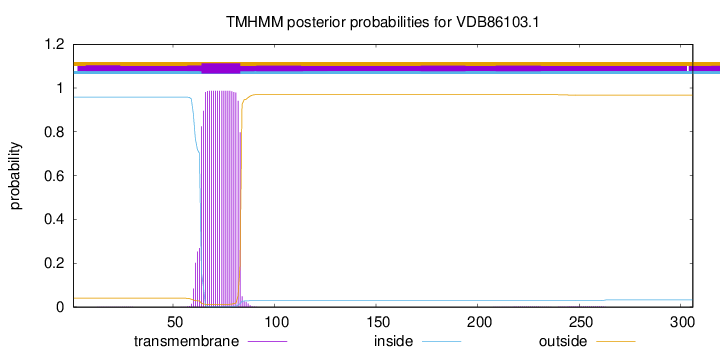
TMHMM annotations
Basic Information help
| Species | Blumeria graminis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Leotiomycetes; ; Erysiphaceae; Blumeria; Blumeria graminis | |||||||||||
| CAZyme ID | VDB86103.1 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE4 | 111 | 232 | 7.3e-25 | 0.9307692307692308 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 200581 | CE4_NodB_like_2 | 2.67e-65 | 113 | 291 | 1 | 188 | Catalytic NodB homology domain of uncharacterized chitin deacetylases and hypothetical proteins. This family includes some uncharacterized chitin deacetylases and hypothetical proteins, mainly from eukaryotes. Although their biological function is unknown, members in this family show high sequence homology to the catalytic NodB homology domain of Colletotrichum lindemuthianum chitin deacetylase (endo-chitin de-N-acetylase, ClCDA, EC 3.5.1.41), which catalyzes the hydrolysis of N-acetamido groups of N-acetyl-D-glucosamine residues in chitin, converting it to chitosan in fungal cell walls. Like ClCDA, this family is a member the carbohydrate esterase 4 (CE4) superfamily. |
| 213022 | CE4_NodB_like_6s_7s | 1.45e-44 | 113 | 280 | 1 | 170 | Catalytic NodB homology domain of rhizobial NodB-like proteins. This family belongs to the large and functionally diverse carbohydrate esterase 4 (CE4) superfamily, whose members show strong sequence similarity with some variability due to their distinct carbohydrate substrates. It includes many rhizobial NodB chitooligosaccharide N-deacetylase (EC 3.5.1.-)-like proteins, mainly from bacteria and eukaryotes, such as chitin deacetylases (EC 3.5.1.41), bacterial peptidoglycan N-acetylglucosamine deacetylases (EC 3.5.1.-), and acetylxylan esterases (EC 3.1.1.72), which catalyze the N- or O-deacetylation of substrates such as acetylated chitin, peptidoglycan, and acetylated xylan. All members of this family contain a catalytic NodB homology domain with the same overall topology and a deformed (beta/alpha)8 barrel fold with 6- or 7 strands. Their catalytic activity is dependent on the presence of a divalent cation, preferably cobalt or zinc, and they employ a conserved His-His-Asp zinc-binding triad closely associated with the conserved catalytic base (aspartic acid) and acid (histidine) to carry out acid/base catalysis. Several family members show diversity both in metal ion specificities and in the residues that coordinate the metal. |
| 200582 | CE4_NodB_like_3 | 4.54e-43 | 113 | 290 | 1 | 187 | Catalytic NodB homology domain of uncharacterized bacterial polysaccharide deacetylases. This family includes many uncharacterized bacterial polysaccharide deacetylases. Although their biological function still remains unknown, members in this family show high sequence homology to the catalytic NodB homology domain of Streptococcus pneumoniae polysaccharide deacetylase PgdA (SpPgdA), which is an extracellular metal-dependent polysaccharide deacetylase with de-N-acetylase activity toward a hexamer of chitooligosaccharide N-acetylglucosamine, but not shorter chitooligosaccharides or a synthetic peptidoglycan tetrasaccharide. Like SpPgdA, this family is a member of the carbohydrate esterase 4 (CE4) superfamily. |
| 274287 | spore_ybaN_pdaB | 1.11e-34 | 113 | 291 | 6 | 188 | polysaccharide deacetylase family sporulation protein PdaB. This model describes the YbaN protein family, also called PdaB and SpoVIE, of Gram-positive bacteria. Although ybaN null mutants have only a mild sporulation defect, ybaN/ytrI double mutants show drastically reducted sporulation efficiencies. This synthetic defect suggests the role of this sigmaE-controlled gene in sporulation had been masked by functional redundancy. Members of this family are homologous to a characterized polysaccharide deacetylase; the exact function this protein family is unknown. [Cellular processes, Sporulation and germination] |
| 223798 | CDA1 | 4.54e-30 | 112 | 297 | 64 | 260 | Peptidoglycan/xylan/chitin deacetylase, PgdA/CDA1 family [Carbohydrate transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.83e-227 | 1 | 306 | 1 | 306 | |
| 3.07e-87 | 33 | 290 | 27 | 294 | |
| 3.25e-87 | 30 | 290 | 24 | 285 | |
| 3.08e-86 | 1 | 290 | 3 | 290 | |
| 3.08e-86 | 1 | 290 | 3 | 290 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.33e-24 | 110 | 291 | 114 | 297 | Chain A, Predicted xylanase/chitin deacetylase [Caldanaerobacter subterraneus subsp. tengcongensis MB4] |
|
| 5.85e-22 | 111 | 298 | 8 | 194 | Structure of peptidoglycan deacetylase PdaC from Bacillus subtilis [Bacillus subtilis subsp. subtilis str. 168],6H8L_B Structure of peptidoglycan deacetylase PdaC from Bacillus subtilis [Bacillus subtilis subsp. subtilis str. 168] |
|
| 4.20e-21 | 111 | 298 | 8 | 194 | Structure of peptidoglycan deacetylase PdaC from Bacillus subtilis - mutant D285S [Bacillus subtilis subsp. subtilis str. 168],6H8N_B Structure of peptidoglycan deacetylase PdaC from Bacillus subtilis - mutant D285S [Bacillus subtilis subsp. subtilis str. 168] |
|
| 8.38e-19 | 111 | 290 | 234 | 412 | Structure of Streptococcus pneumoniae peptidoglycan deacetylase (SpPgdA) [Streptococcus pneumoniae R6] |
|
| 3.85e-18 | 111 | 290 | 234 | 412 | Structure of Streptococcus pneumoniae peptidoglycan deacetylase (SpPgdA) D 275 N Mutant. [Streptococcus pneumoniae R6] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.07e-20 | 111 | 298 | 276 | 462 | Peptidoglycan-N-acetylmuramic acid deacetylase PdaC OS=Bacillus subtilis (strain 168) OX=224308 GN=pdaC PE=1 SV=1 |
|
| 2.96e-18 | 110 | 264 | 19 | 176 | Peptidoglycan-N-acetylglucosamine deacetylase BC_3618 OS=Bacillus cereus (strain ATCC 14579 / DSM 31 / CCUG 7414 / JCM 2152 / NBRC 15305 / NCIMB 9373 / NCTC 2599 / NRRL B-3711) OX=226900 GN=BC_3618 PE=1 SV=1 |
|
| 4.94e-18 | 111 | 290 | 266 | 444 | Peptidoglycan-N-acetylglucosamine deacetylase OS=Streptococcus pneumoniae (strain ATCC BAA-255 / R6) OX=171101 GN=pgdA PE=1 SV=1 |
|
| 8.82e-18 | 112 | 267 | 20 | 178 | Chitooligosaccharide deacetylase OS=Mesorhizobium japonicum (strain LMG 29417 / CECT 9101 / MAFF 303099) OX=266835 GN=nodB PE=3 SV=2 |
|
| 4.14e-17 | 107 | 290 | 260 | 442 | Peptidoglycan-N-acetylglucosamine deacetylase PgdA OS=Listeria monocytogenes serovar 1/2a (strain ATCC BAA-679 / EGD-e) OX=169963 GN=pgdA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999223 | 0.000788 |