You are browsing environment: FUNGIDB
CAZyme Information: TSTA_097140-t26_1-p1
You are here: Home > Sequence: TSTA_097140-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Talaromyces stipitatus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Trichocomaceae; Talaromyces; Talaromyces stipitatus | |||||||||||
| CAZyme ID | TSTA_097140-t26_1-p1 | |||||||||||
| CAZy Family | GH55|GH55 | |||||||||||
| CAZyme Description | extracellular polygalacturonase, putative | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.15:80 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 57 | 383 | 2.9e-71 | 0.963076923076923 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395231 | Glyco_hydro_28 | 1.22e-115 | 60 | 382 | 1 | 321 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| 178580 | PLN03003 | 9.30e-27 | 111 | 341 | 114 | 336 | Probable polygalacturonase At3g15720 |
| 177865 | PLN02218 | 1.91e-21 | 57 | 375 | 92 | 417 | polygalacturonase ADPG |
| 227721 | Pgu1 | 3.51e-21 | 155 | 297 | 251 | 398 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| 165802 | PLN02155 | 3.22e-19 | 144 | 368 | 150 | 367 | polygalacturonase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 6.12e-231 | 10 | 383 | 10 | 383 | |
| 8.56e-211 | 9 | 383 | 12 | 386 | |
| 1.27e-201 | 14 | 383 | 13 | 380 | |
| 1.99e-191 | 13 | 383 | 12 | 379 | |
| 1.99e-191 | 13 | 383 | 12 | 379 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.85e-145 | 39 | 382 | 1 | 339 | Polygalacturonase From Aspergillus Aculeatus [Aspergillus aculeatus],1IB4_A Crystal Structure of Polygalacturonase from Aspergillus Aculeatus at Ph4.5 [Aspergillus aculeatus],1IB4_B Crystal Structure of Polygalacturonase from Aspergillus Aculeatus at Ph4.5 [Aspergillus aculeatus] |
|
| 1.45e-140 | 31 | 382 | 1 | 344 | Chain A, Endo-polygalacturonase [Evansstolkia leycettana] |
|
| 1.45e-140 | 31 | 382 | 1 | 344 | Chain A, endo-polygalacturonase [Evansstolkia leycettana] |
|
| 2.07e-138 | 41 | 382 | 2 | 336 | Chain A, Endo-polygalacturonase [Evansstolkia leycettana] |
|
| 2.15e-136 | 41 | 378 | 2 | 333 | Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_B Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_C Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_D Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_E Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_F Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_G Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.54e-192 | 13 | 383 | 12 | 379 | Endopolygalacturonase AN8327 OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=AN8327 PE=1 SV=1 |
|
| 5.42e-191 | 10 | 382 | 9 | 378 | Probable endopolygalacturonase AFUB_016610 OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=AFUB_016610 PE=3 SV=1 |
|
| 5.42e-191 | 10 | 382 | 9 | 378 | Probable endopolygalacturonase AFUA_1G17220 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=AFUA_1G17220 PE=3 SV=1 |
|
| 5.97e-188 | 10 | 382 | 9 | 378 | Probable endopolygalacturonase NFIA_008150 OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=NFIA_008150 PE=3 SV=1 |
|
| 7.38e-158 | 20 | 382 | 19 | 378 | Probable endopolygalacturonase E OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=pgaE PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
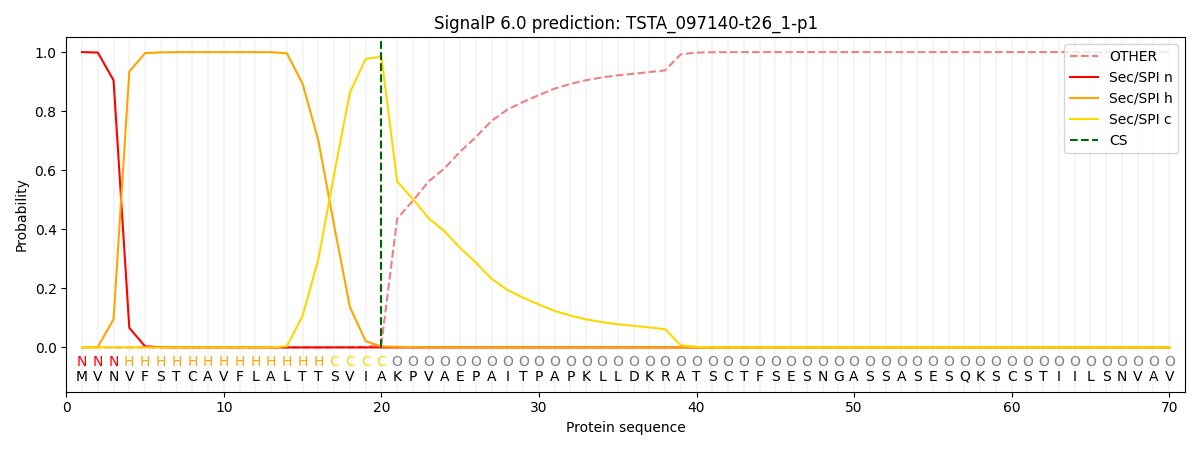
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000236 | 0.999731 | CS pos: 20-21. Pr: 0.9841 |
