You are browsing environment: FUNGIDB
CAZyme Information: TSTA_069640-t26_1-p1
You are here: Home > Sequence: TSTA_069640-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Talaromyces stipitatus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Trichocomaceae; Talaromyces; Talaromyces stipitatus | |||||||||||
| CAZyme ID | TSTA_069640-t26_1-p1 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | extracellular endoglucanase, putative | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.-:12 | 3.2.1.4:4 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 56 | 318 | 4.6e-60 | 0.9891304347826086 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395098 | Cellulase | 2.49e-28 | 56 | 320 | 16 | 271 | Cellulase (glycosyl hydrolase family 5). |
| 225344 | BglC | 1.89e-15 | 34 | 316 | 45 | 358 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| 397483 | CBM_X2 | 8.18e-12 | 363 | 445 | 3 | 77 | Carbohydrate binding domain X2. This domain binds to cellulose and to bacterial cell walls. It is found in glycosyl hydrolases and in scaffolding proteins of cellulosomes (multiprotein glycosyl hydrolase complexes). In the cellulosome it may aid cellulose degradation by anchoring the cellulosome to the bacterial cell wall and by binding it to its substrate. This domain has an Ig-like fold. |
| 176230 | Zn_ADH9 | 0.005 | 165 | 248 | 175 | 243 | Alcohol dehydrogenases of the MDR family. The medium chain dehydrogenases/reductase (MDR)/zinc-dependent alcohol dehydrogenase-like family, which contains the zinc-dependent alcohol dehydrogenase (ADH-Zn) and related proteins, is a diverse group of proteins related to the first identified member, class I mammalian ADH. MDRs display a broad range of activities and are distinguished from the smaller short chain dehydrogenases (~ 250 amino acids vs. the ~ 350 amino acids of the MDR). The MDR proteins have 2 domains: a C-terminal NAD(P)-binding Rossmann fold domain of a beta-alpha form and an N-terminal catalytic domain with distant homology to GroES. The MDR group contains a host of activities, including the founding alcohol dehydrogenase (ADH), quinone reductase, sorbitol dehydrogenase, formaldehyde dehydrogenase, butanediol DH, ketose reductase, cinnamyl reductase, and numerous others. The zinc-dependent alcohol dehydrogenases (ADHs) catalyze the NAD(P)(H)-dependent interconversion of alcohols to aldehydes or ketones. Active site zinc has a catalytic role, while structural zinc aids in stability. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.77e-197 | 13 | 569 | 9 | 574 | |
| 1.42e-196 | 3 | 569 | 2 | 572 | |
| 2.57e-196 | 9 | 569 | 7 | 569 | |
| 2.57e-196 | 9 | 569 | 7 | 569 | |
| 1.04e-195 | 9 | 569 | 7 | 569 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.58e-79 | 31 | 554 | 22 | 524 | Crystal structure of a tri-modular GH5 (subfamily 4) endo-beta-1, 4-glucanase from Bacillus licheniformis [Bacillus licheniformis],4YZT_A Crystal structure of a tri-modular GH5 (subfamily 4) endo-beta-1, 4-glucanase from Bacillus licheniformis complexed with cellotetraose [Bacillus licheniformis] |
|
| 2.24e-46 | 36 | 504 | 13 | 477 | Structural Insight of a Trimodular Halophilic Cellulase with a Family 46 Carbohydrate-Binding Module [Bacillus sp. BG-CS10] |
|
| 3.97e-46 | 4 | 547 | 28 | 562 | Chain A, Endo-beta-1,4-glucanase (cellulase B) [Halalkalibacterium halodurans] |
|
| 4.62e-46 | 36 | 504 | 46 | 510 | A Trimodular GH5_4 Subfamily Endoglucanase Structure with Large Unit Cell [Bacillus sp. BG-CS10],5XRC_B A Trimodular GH5_4 Subfamily Endoglucanase Structure with Large Unit Cell [Bacillus sp. BG-CS10],5XRC_C A Trimodular GH5_4 Subfamily Endoglucanase Structure with Large Unit Cell [Bacillus sp. BG-CS10] |
|
| 2.88e-45 | 36 | 504 | 13 | 477 | Structural Insight of a Trimodular Halophilic Cellulase with a Family 46 Carbohydrate-Binding Module [Bacillus sp. BG-CS10] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.84e-50 | 3 | 496 | 7 | 496 | Endoglucanase B OS=Paenibacillus lautus OX=1401 GN=celB PE=3 SV=1 |
|
| 5.57e-41 | 31 | 348 | 41 | 378 | Endoglucanase D OS=Clostridium cellulovorans (strain ATCC 35296 / DSM 3052 / OCM 3 / 743B) OX=573061 GN=engD PE=1 SV=2 |
|
| 1.03e-39 | 31 | 342 | 55 | 385 | Cellulase/esterase CelE OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celE PE=1 SV=2 |
|
| 1.36e-39 | 31 | 356 | 40 | 392 | Endoglucanase B OS=Clostridium cellulovorans (strain ATCC 35296 / DSM 3052 / OCM 3 / 743B) OX=573061 GN=engB PE=3 SV=1 |
|
| 1.52e-36 | 28 | 341 | 23 | 331 | Endoglucanase D OS=Ruminiclostridium cellulolyticum (strain ATCC 35319 / DSM 5812 / JCM 6584 / H10) OX=394503 GN=celCCD PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
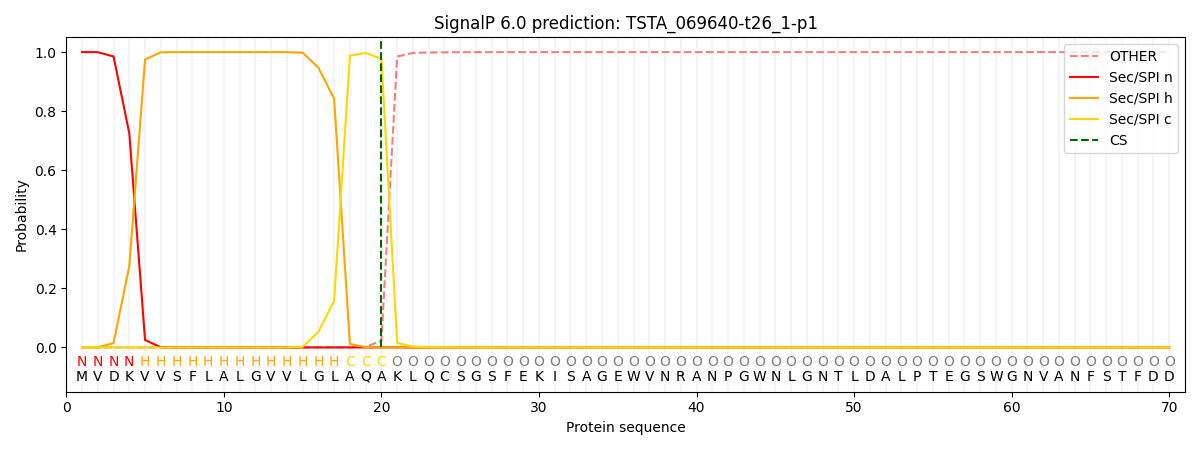
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000197 | 0.999778 | CS pos: 20-21. Pr: 0.9775 |
