You are browsing environment: FUNGIDB
CAZyme Information: TSTA_033310-t26_1-p1
You are here: Home > Sequence: TSTA_033310-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
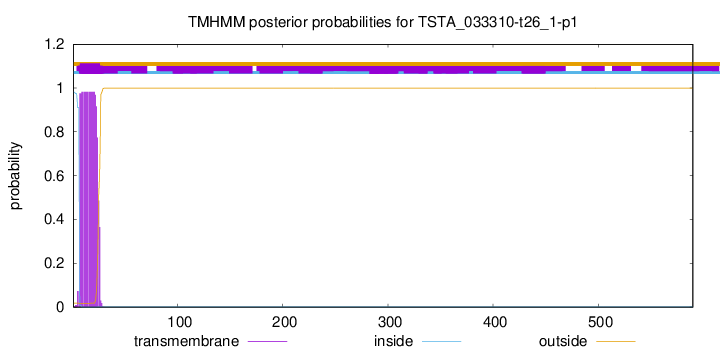
TMHMM annotations
Basic Information help
| Species | Talaromyces stipitatus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Trichocomaceae; Talaromyces; Talaromyces stipitatus | |||||||||||
| CAZyme ID | TSTA_033310-t26_1-p1 | |||||||||||
| CAZy Family | CE8 | |||||||||||
| CAZyme Description | capsule-associated protein CAP1, putative | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 214773 | CAP10 | 7.50e-17 | 335 | 578 | 32 | 252 | Putative lipopolysaccharide-modifying enzyme. |
| 310354 | Glyco_transf_90 | 3.03e-10 | 335 | 578 | 108 | 322 | Glycosyl transferase family 90. This family of glycosyl transferases are specifically (mannosyl) glucuronoxylomannan/galactoxylomannan -beta 1,2-xylosyltransferases, EC:2.4.2.-. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 590 | 1 | 590 | |
| 0.0 | 1 | 588 | 298 | 886 | |
| 2.08e-291 | 1 | 589 | 1 | 599 | |
| 2.09e-279 | 1 | 588 | 11 | 617 | |
| 1.69e-278 | 1 | 588 | 1 | 607 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.81e-07 | 312 | 579 | 95 | 342 | human POGLUT1 in complex with Notch1 EGF12 and UDP [Homo sapiens],5L0S_A human POGLUT1 in complex with Factor VII EGF1 and UDP [Homo sapiens],5L0T_A human POGLUT1 in complex with EGF(+) and UDP [Homo sapiens],5L0U_A human POGLUT1 in complex with EGF(+) and UDP-phosphono-glucose [Homo sapiens],5L0V_A human POGLUT1 in complex with 2F-glucose modified EGF(+) and UDP [Homo sapiens],5UB5_A human POGLUT1 in complex with human Notch1 EGF12 S458T mutant and UDP [Homo sapiens] |
|
| 1.00e-06 | 335 | 579 | 154 | 379 | Crystal structure of Drosophila Poglut1 (Rumi) complexed with its glycoprotein product (glucosylated EGF repeat) and UDP [Drosophila melanogaster],5F85_A Crystal structure of Drosophila Poglut1 (Rumi) complexed with its substrate protein (EGF repeat) and UDP [Drosophila melanogaster],5F86_A Crystal structure of Drosophila Poglut1 (Rumi) complexed with its substrate protein (EGF repeat) [Drosophila melanogaster],5F87_A Crystal structure of Drosophila Poglut1 (Rumi) complexed with UDP [Drosophila melanogaster],5F87_B Crystal structure of Drosophila Poglut1 (Rumi) complexed with UDP [Drosophila melanogaster],5F87_C Crystal structure of Drosophila Poglut1 (Rumi) complexed with UDP [Drosophila melanogaster],5F87_D Crystal structure of Drosophila Poglut1 (Rumi) complexed with UDP [Drosophila melanogaster],5F87_E Crystal structure of Drosophila Poglut1 (Rumi) complexed with UDP [Drosophila melanogaster],5F87_F Crystal structure of Drosophila Poglut1 (Rumi) complexed with UDP [Drosophila melanogaster] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.19e-59 | 55 | 585 | 151 | 662 | Beta-1,2-xylosyltransferase 1 OS=Cryptococcus neoformans var. neoformans serotype D (strain JEC21 / ATCC MYA-565) OX=214684 GN=CXT1 PE=1 SV=1 |
|
| 1.10e-09 | 328 | 579 | 157 | 389 | O-glucosyltransferase rumi homolog OS=Culex quinquefasciatus OX=7176 GN=CPIJ013394 PE=3 SV=1 |
|
| 2.57e-09 | 331 | 579 | 159 | 388 | O-glucosyltransferase rumi homolog OS=Aedes aegypti OX=7159 GN=AAEL011121 PE=3 SV=1 |
|
| 1.72e-07 | 312 | 579 | 123 | 370 | Protein O-glucosyltransferase 1 OS=Rattus norvegicus OX=10116 GN=Poglut1 PE=3 SV=1 |
|
| 2.87e-06 | 312 | 579 | 123 | 370 | Protein O-glucosyltransferase 1 OS=Mus musculus OX=10090 GN=Poglut1 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
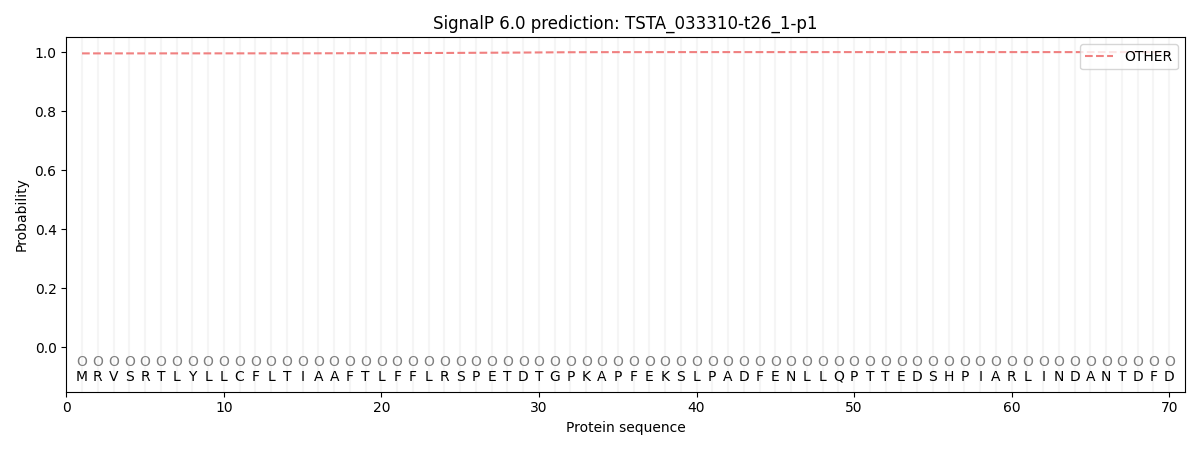
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.995946 | 0.004080 |