You are browsing environment: FUNGIDB
CAZyme Information: TSTA_010240-t26_1-p1
You are here: Home > Sequence: TSTA_010240-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Talaromyces stipitatus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Trichocomaceae; Talaromyces; Talaromyces stipitatus | |||||||||||
| CAZyme ID | TSTA_010240-t26_1-p1 | |||||||||||
| CAZy Family | AA3 | |||||||||||
| CAZyme Description | glycosyl hydrolase, putative | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 86 | 311 | 1.4e-44 | 0.9675925925925926 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395747 | Glyco_hydro_3 | 6.84e-63 | 42 | 351 | 19 | 316 | Glycosyl hydrolase family 3 N terminal domain. |
| 224389 | BglX | 3.22e-53 | 44 | 365 | 19 | 325 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| 235417 | PRK05337 | 7.18e-37 | 55 | 353 | 26 | 307 | beta-hexosaminidase; Provisional |
| 185053 | PRK15098 | 7.33e-07 | 119 | 349 | 129 | 350 | beta-glucosidase BglX. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.45e-202 | 1 | 353 | 1 | 353 | |
| 1.38e-167 | 30 | 353 | 24 | 347 | |
| 2.27e-131 | 17 | 353 | 19 | 355 | |
| 1.61e-128 | 29 | 348 | 27 | 348 | |
| 2.22e-126 | 29 | 347 | 23 | 341 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.80e-36 | 25 | 352 | 13 | 337 | Structure of a glycoside hydrolase family 3 beta-N-acetylglucosaminidase from Paenibacillus sp. str. FPU-7 [Paenibacillaceae] |
|
| 9.71e-29 | 25 | 314 | 7 | 300 | Crystal structure of Beta-N-acetylhexosaminidase from Beutenbergia cavernae DSM 12333 [Beutenbergia cavernae DSM 12333],5BU9_B Crystal structure of Beta-N-acetylhexosaminidase from Beutenbergia cavernae DSM 12333 [Beutenbergia cavernae DSM 12333] |
|
| 4.11e-28 | 21 | 352 | 5 | 337 | A unique GCN5-related glucosamine N-acetyltransferase region exist in the fungal multi-domain GH3 beta-N-acetylglucosaminidase [Rhizomucor miehei CAU432],4ZM6_B A unique GCN5-related glucosamine N-acetyltransferase region exist in the fungal multi-domain GH3 beta-N-acetylglucosaminidase [Rhizomucor miehei CAU432] |
|
| 1.04e-26 | 22 | 348 | 19 | 342 | Crystal Structure of Glycoside Hydrolase from Synechococcus [Synechococcus sp. PCC 7002],3SQL_B Crystal Structure of Glycoside Hydrolase from Synechococcus [Synechococcus sp. PCC 7002],3SQM_A Crystal Structure of Glycoside Hydrolase from Synechococcus Complexed with N-acetyl-D-glucosamine [Synechococcus sp. PCC 7002],3SQM_B Crystal Structure of Glycoside Hydrolase from Synechococcus Complexed with N-acetyl-D-glucosamine [Synechococcus sp. PCC 7002],3SQM_C Crystal Structure of Glycoside Hydrolase from Synechococcus Complexed with N-acetyl-D-glucosamine [Synechococcus sp. PCC 7002],3SQM_D Crystal Structure of Glycoside Hydrolase from Synechococcus Complexed with N-acetyl-D-glucosamine [Synechococcus sp. PCC 7002] |
|
| 2.74e-26 | 30 | 309 | 18 | 291 | Crystal structure of NagZ H174A mutant from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G5U_B Crystal structure of NagZ H174A mutant from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G6T_A Crystal structure of Zn-containing NagZ H174A mutant from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G6T_B Crystal structure of Zn-containing NagZ H174A mutant from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5LY7_A Crystal structure of NagZ H174A mutant from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1],5LY7_B Crystal structure of NagZ H174A mutant from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.04e-123 | 30 | 351 | 26 | 347 | Uncharacterized secreted glycosyl hydrolase ARB_06359 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_06359 PE=1 SV=1 |
|
| 6.99e-33 | 45 | 345 | 16 | 304 | Beta-hexosaminidase OS=Tolumonas auensis (strain DSM 9187 / TA4) OX=595494 GN=nagZ PE=3 SV=1 |
|
| 4.37e-27 | 50 | 336 | 19 | 309 | Beta-hexosaminidase OS=Stenotrophomonas maltophilia (strain K279a) OX=522373 GN=nagZ PE=3 SV=1 |
|
| 5.92e-27 | 50 | 336 | 19 | 309 | Beta-hexosaminidase OS=Xanthomonas axonopodis pv. citri (strain 306) OX=190486 GN=nagZ PE=3 SV=1 |
|
| 8.31e-27 | 50 | 336 | 19 | 309 | Beta-hexosaminidase OS=Stenotrophomonas maltophilia (strain R551-3) OX=391008 GN=nagZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
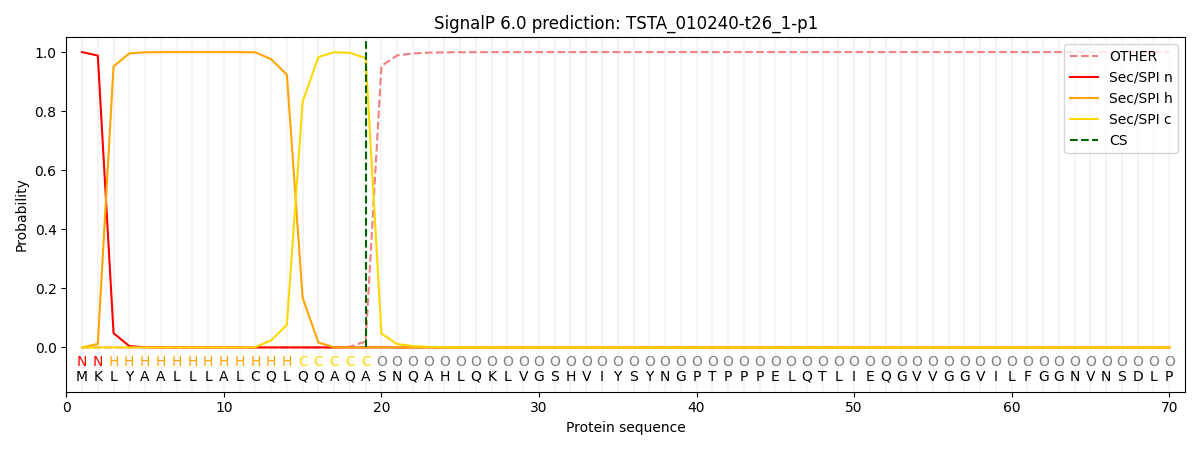
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000199 | 0.999768 | CS pos: 19-20. Pr: 0.9798 |
