You are browsing environment: FUNGIDB
CAZyme Information: TRIVIDRAFT_32988-t45_1-p1
You are here: Home > Sequence: TRIVIDRAFT_32988-t45_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Trichoderma virens | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Hypocreaceae; Trichoderma; Trichoderma virens | |||||||||||
| CAZyme ID | TRIVIDRAFT_32988-t45_1-p1 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 4.2.2.14:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL20 | 23 | 258 | 7.8e-101 | 0.9956331877729258 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 404905 | Polysacc_lyase | 1.42e-72 | 25 | 245 | 2 | 213 | Polysaccharide lyase. This family includes heparin lyase I, EC:4.2.2.7. Heparin lyase I depolymerizes heparin by cleaving the glycosidic linkage next to an iduronic acid moiety. The structure of heparin lyase I consists of a beta-jelly roll domain with a long, deep substrate-binding groove and an unusual thumb domain containing many basic residues extending from the main body of the enzyme. This family also includes glucuronan lyase, EC:4.2.2.14. The structure glucuronan lyase is a beta-jelly roll. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 6.00e-198 | 1 | 259 | 1 | 259 | |
| 6.00e-198 | 1 | 259 | 1 | 259 | |
| 6.95e-188 | 1 | 259 | 1 | 259 | |
| 3.99e-180 | 1 | 259 | 1 | 258 | |
| 2.37e-172 | 1 | 259 | 1 | 258 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.15e-125 | 20 | 257 | 1 | 237 | Chain A, Glucuronan lyase A [Trichoderma reesei] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
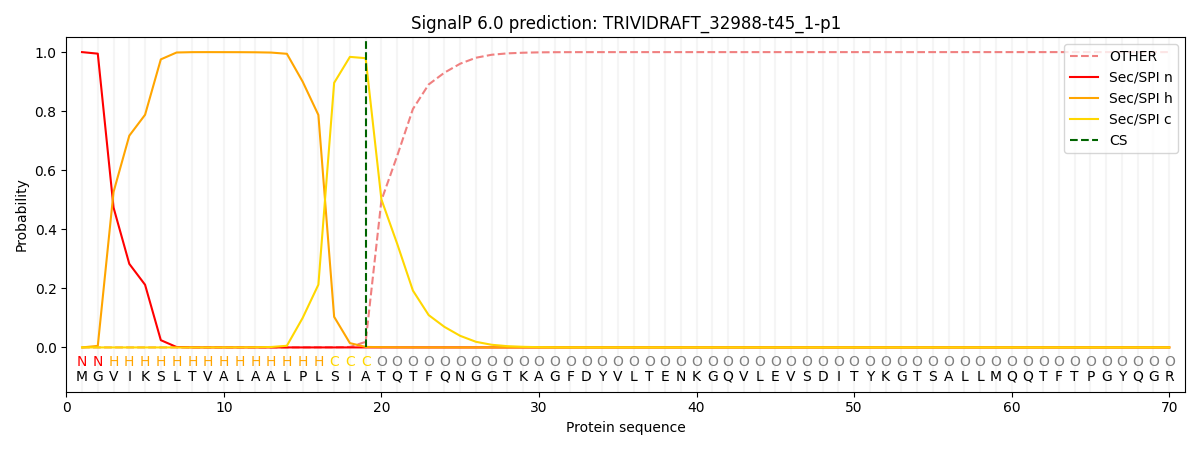
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000337 | 0.999626 | CS pos: 19-20. Pr: 0.9794 |
