You are browsing environment: FUNGIDB
CAZyme Information: TRIREDRAFT_75015-t26_1-p1
You are here: Home > Sequence: TRIREDRAFT_75015-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Trichoderma reesei | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Hypocreaceae; Trichoderma; Trichoderma reesei | |||||||||||
| CAZyme ID | TRIREDRAFT_75015-t26_1-p1 | |||||||||||
| CAZy Family | GT22 | |||||||||||
| CAZyme Description | glycoside hydrolase family 27 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.22:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH27 | 184 | 453 | 4.5e-41 | 0.9519650655021834 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 269893 | GH27 | 8.15e-66 | 50 | 365 | 2 | 268 | glycosyl hydrolase family 27 (GH27). GH27 enzymes occur in eukaryotes, prokaryotes, and archaea with a wide range of hydrolytic activities, including alpha-glucosidase (glucoamylase and sucrase-isomaltase), alpha-N-acetylgalactosaminidase, and 3-alpha-isomalto-dextranase. All GH27 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. GH27 members are retaining enzymes that cleave their substrates via an acid/base-catalyzed, double-displacement mechanism involving a covalent glycosyl-enzyme intermediate. Two aspartic acid residues have been identified as the catalytic nucleophile and the acid/base, respectively. |
| 177874 | PLN02229 | 4.07e-15 | 5 | 458 | 13 | 401 | alpha-galactosidase |
| 166449 | PLN02808 | 1.37e-12 | 52 | 475 | 35 | 385 | alpha-galactosidase |
| 178295 | PLN02692 | 5.58e-11 | 52 | 458 | 59 | 392 | alpha-galactosidase |
| 374582 | Melibiase_2 | 9.52e-11 | 87 | 360 | 47 | 276 | Alpha galactosidase A. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.12e-314 | 1 | 476 | 1 | 467 | |
| 5.23e-311 | 1 | 475 | 1 | 476 | |
| 1.29e-284 | 62 | 476 | 19 | 433 | |
| 6.14e-237 | 9 | 476 | 7 | 473 | |
| 1.24e-236 | 9 | 476 | 7 | 473 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.62e-22 | 52 | 476 | 28 | 443 | Crystal structure of Abp-WT, a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NX0_B Crystal structure of Abp-WT, a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NX0_C Crystal structure of Abp-WT, a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NX0_D Crystal structure of Abp-WT, a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NX0_E Crystal structure of Abp-WT, a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NX0_F Crystal structure of Abp-WT, a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NX0_G Crystal structure of Abp-WT, a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NX0_H Crystal structure of Abp-WT, a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus] |
|
| 2.81e-21 | 52 | 476 | 28 | 443 | Crystal structure of Abp-D197A, a catalytic mutant of a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NXK_B Crystal structure of Abp-D197A, a catalytic mutant of a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NXK_C Crystal structure of Abp-D197A, a catalytic mutant of a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NXK_D Crystal structure of Abp-D197A, a catalytic mutant of a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NXK_E Crystal structure of Abp-D197A, a catalytic mutant of a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NXK_F Crystal structure of Abp-D197A, a catalytic mutant of a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NXK_G Crystal structure of Abp-D197A, a catalytic mutant of a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NXK_H Crystal structure of Abp-D197A, a catalytic mutant of a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4NZF_A Crystal structure of Abp-D197A (a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus), in complex with arabinose [Geobacillus stearothermophilus],4NZF_B Crystal structure of Abp-D197A (a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus), in complex with arabinose [Geobacillus stearothermophilus],4NZF_C Crystal structure of Abp-D197A (a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus), in complex with arabinose [Geobacillus stearothermophilus],4NZF_D Crystal structure of Abp-D197A (a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus), in complex with arabinose [Geobacillus stearothermophilus],4NZF_E Crystal structure of Abp-D197A (a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus), in complex with arabinose [Geobacillus stearothermophilus],4NZF_F Crystal structure of Abp-D197A (a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus), in complex with arabinose [Geobacillus stearothermophilus],4NZF_G Crystal structure of Abp-D197A (a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus), in complex with arabinose [Geobacillus stearothermophilus],4NZF_H Crystal structure of Abp-D197A (a GH27-b-L-arabinopyranosidase from Geobacillus stearothermophilus), in complex with arabinose [Geobacillus stearothermophilus] |
|
| 1.23e-12 | 47 | 474 | 7 | 361 | Nicotiana benthamiana alpha-galactosidase [Nicotiana benthamiana] |
|
| 4.54e-10 | 53 | 359 | 16 | 282 | Chain A, Alpha-N-acetylgalactosaminidase [Homo sapiens],3H53_B Chain B, Alpha-N-acetylgalactosaminidase [Homo sapiens],3H54_A Chain A, Alpha-N-acetylgalactosaminidase [Homo sapiens],3H54_B Chain B, Alpha-N-acetylgalactosaminidase [Homo sapiens],3H55_A Chain A, Alpha-N-acetylgalactosaminidase [Homo sapiens],3H55_B Chain B, Alpha-N-acetylgalactosaminidase [Homo sapiens],3IGU_A Chain A, Alpha-N-acetylgalactosaminidase [Homo sapiens],3IGU_B Chain B, Alpha-N-acetylgalactosaminidase [Homo sapiens],4DO4_A Pharmacological chaperones for human alpha-N-acetylgalactosaminidase [Homo sapiens],4DO4_B Pharmacological chaperones for human alpha-N-acetylgalactosaminidase [Homo sapiens],4DO5_A Pharmacological chaperones for human alpha-N-acetylgalactosaminidase [Homo sapiens],4DO5_B Pharmacological chaperones for human alpha-N-acetylgalactosaminidase [Homo sapiens],4DO6_A Pharmacological chaperones for human alpha-N-acetylgalactosaminidase [Homo sapiens],4DO6_B Pharmacological chaperones for human alpha-N-acetylgalactosaminidase [Homo sapiens] |
|
| 6.67e-10 | 104 | 474 | 75 | 430 | Chain A, Putative alpha-N-acetylgalactosaminidase [Halalkalibacterium halodurans C-125],3CC1_B Chain B, Putative alpha-N-acetylgalactosaminidase [Halalkalibacterium halodurans C-125] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.84e-139 | 34 | 475 | 168 | 622 | Alpha-galactosidase 3 OS=Hypocrea jecorina OX=51453 GN=agl3 PE=1 SV=1 |
|
| 2.08e-93 | 18 | 436 | 56 | 477 | Putative alpha-galactosidase 8 OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=agl8 PE=5 SV=2 |
|
| 8.73e-14 | 37 | 476 | 1 | 425 | Probable alpha-galactosidase B OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=aglB PE=3 SV=1 |
|
| 1.27e-13 | 50 | 476 | 34 | 446 | Probable alpha-galactosidase B OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=aglB PE=3 SV=2 |
|
| 1.63e-13 | 47 | 458 | 71 | 411 | Alpha-galactosidase 3 OS=Arabidopsis thaliana OX=3702 GN=AGAL3 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
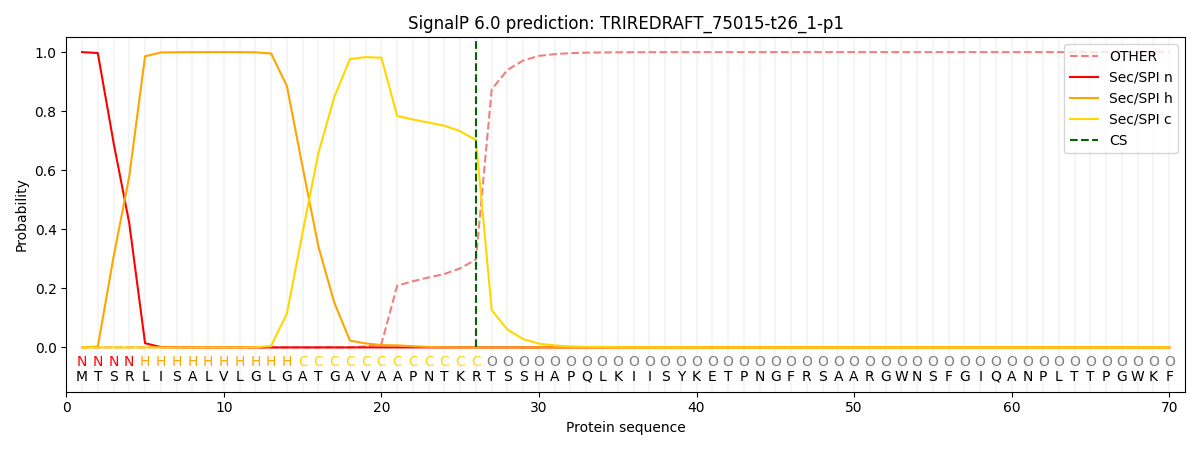
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000385 | 0.999567 | CS pos: 26-27. Pr: 0.7023 |
