You are browsing environment: FUNGIDB
CAZyme Information: THC96494.1
You are here: Home > Sequence: THC96494.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus tanneri | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus tanneri | |||||||||||
| CAZyme ID | THC96494.1 | |||||||||||
| CAZy Family | GH71 | |||||||||||
| CAZyme Description | 1,4-alpha-D-glucan glucohydrolase [Source:UniProtKB/TrEMBL;Acc:A0A4V3UPU3] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.3:94 | 1.14.99.55:4 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH15 | 48 | 459 | 4.2e-77 | 0.96398891966759 |
| CBM20 | 460 | 537 | 5.8e-18 | 0.8444444444444444 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395586 | Glyco_hydro_15 | 9.17e-114 | 44 | 459 | 3 | 408 | Glycosyl hydrolases family 15. In higher organisms this family is represented by phosphorylase kinase subunits. |
| 395557 | CBM_20 | 6.06e-22 | 461 | 540 | 13 | 93 | Starch binding domain. |
| 99886 | CBM20_glucoamylase | 3.55e-19 | 462 | 545 | 20 | 106 | Glucoamylase (glucan1,4-alpha-glucosidase), C-terminal CBM20 (carbohydrate-binding module, family 20) domain. Glucoamylases are inverting, exo-acting starch hydrolases that hydrolyze starch and related polysaccharides by releasing the nonreducing end glucose. They are mainly active on alpha-1,4-glycosidic bonds but also have some activity towards 1,6-glycosidic bonds occurring in natural oligosaccharides. The ability of glucoamylases to cleave 1-6-glycosidic binds is called "debranching activity" and is of importance in industrial applications, where complete degradation of starch to glucose is needed. Most glucoamylases are multidomain proteins containing an N-terminal catalytic domain, a C-terminal CBM20 domain, and a highly O-glycosylated linker region that connects the two. The CBM20 domain is found in a large number of starch degrading enzymes including alpha-amylase, beta-amylase, glucoamylase, and CGTase (cyclodextrin glucanotransferase). CBM20 is also present in proteins that have a regulatory role in starch metabolism in plants (e.g. alpha-amylase) or glycogen metabolism in mammals (e.g. laforin). CBM20 folds as an antiparallel beta-barrel structure with two starch binding sites. These two sites are thought to differ functionally with site 1 acting as the initial starch recognition site and site 2 involved in the specific recognition of appropriate regions of starch. |
| 99883 | CBM20_alpha_amylase | 8.60e-16 | 451 | 546 | 7 | 95 | Alpha-amylase, C-terminal CBM20 (carbohydrate-binding module, family 20) domain. This domain is found in several bacterial and fungal alpha-amylases including the maltopentaose-forming amylases (G5-amylases). Most alpha-amylases have, in addition to the C-terminal CBM20 domain, an N-terminal catalytic domain belonging to glycosyl hydrolase family 13, which hydrolyzes internal alpha-1,4-glucosidic bonds in starch and related saccharides, yielding maltotriose and maltose. Two types of soluble substrates are used by alpha-amylases including long substrates (e.g. amylose) and short substrates (e.g. maltodextrins or maltooligosaccharides). The CBM20 domain is found in a large number of starch degrading enzymes including alpha-amylase, beta-amylase, glucoamylase, and CGTase (cyclodextrin glucanotransferase). CBM20 is also present in proteins that have a regulatory role in starch metabolism in plants (e.g. alpha-amylase) or glycogen metabolism in mammals (e.g. laforin). CBM20 folds as an antiparallel beta-barrel structure with two starch binding sites. These two sites are thought to differ functionally with site 1 acting as the initial starch recognition site and site 2 involved in the specific recognition of appropriate regions of starch. |
| 119437 | CBM20 | 1.20e-15 | 461 | 545 | 12 | 96 | The family 20 carbohydrate-binding module (CBM20), also known as the starch-binding domain, is found in a large number of starch degrading enzymes including alpha-amylase, beta-amylase, glucoamylase, and CGTase (cyclodextrin glucanotransferase). CBM20 is also present in proteins that have a regulatory role in starch metabolism in plants (e.g. alpha-amylase) or glycogen metabolism in mammals (e.g. laforin). CBM20 folds as an antiparallel beta-barrel structure with two starch binding sites. These two sites are thought to differ functionally with site 1 acting as the initial starch recognition site and site 2 involved in the specific recognition of appropriate regions of starch. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 9.73e-190 | 28 | 536 | 27 | 606 | |
| 9.73e-190 | 28 | 536 | 27 | 606 | |
| 9.73e-190 | 28 | 536 | 27 | 606 | |
| 9.73e-190 | 28 | 536 | 27 | 606 | |
| 4.51e-189 | 32 | 536 | 4 | 579 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.07e-167 | 32 | 546 | 7 | 593 | Crystal structure of Penicillium oxalicum Glucoamylase [Penicillium oxalicum 114-2] |
|
| 1.48e-165 | 32 | 546 | 2 | 616 | Structure of the catalytic domain of Aspergillus niger Glucoamylase [Aspergillus niger] |
|
| 1.31e-161 | 32 | 459 | 2 | 417 | Refined structure for the complex of acarbose with glucoamylase from Aspergillus awamori var. x100 to 2.4 angstroms resolution [Aspergillus awamori],1DOG_A REFINED STRUCTURE FOR THE COMPLEX OF 1-DEOXYNOJIRIMYCIN WITH GLUCOAMYLASE FROM (ASPERGILLUS AWAMORI) VAR. X100 TO 2.4 ANGSTROMS RESOLUTION [Aspergillus awamori],1GLM_A REFINED CRYSTAL STRUCTURES OF GLUCOAMYLASE FROM ASPERGILLUS AWAMORI VAR. X100 [Aspergillus awamori],3GLY_A REFINED CRYSTAL STRUCTURES OF GLUCOAMYLASE FROM ASPERGILLUS AWAMORI VAR. X100 [Aspergillus awamori] |
|
| 1.35e-161 | 32 | 459 | 2 | 417 | GLUCOAMYLASE-471 COMPLEXED WITH ACARBOSE [Aspergillus awamori] |
|
| 1.40e-161 | 32 | 459 | 2 | 417 | GLUCOAMYLASE-471 COMPLEXED WITH D-GLUCO-DIHYDROACARBOSE [Aspergillus awamori] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.88e-167 | 28 | 536 | 22 | 598 | Glucoamylase ARB_02327-1 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_02327-1 PE=1 SV=1 |
|
| 4.74e-165 | 28 | 546 | 25 | 612 | Glucoamylase OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=glaA PE=2 SV=2 |
|
| 8.17e-165 | 23 | 546 | 17 | 640 | Glucoamylase OS=Aspergillus awamori OX=105351 GN=GLAA PE=1 SV=1 |
|
| 8.17e-165 | 23 | 546 | 17 | 640 | Glucoamylase OS=Aspergillus niger OX=5061 GN=GLAA PE=1 SV=1 |
|
| 4.49e-164 | 23 | 546 | 17 | 639 | Glucoamylase OS=Aspergillus usamii OX=186680 GN=glaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
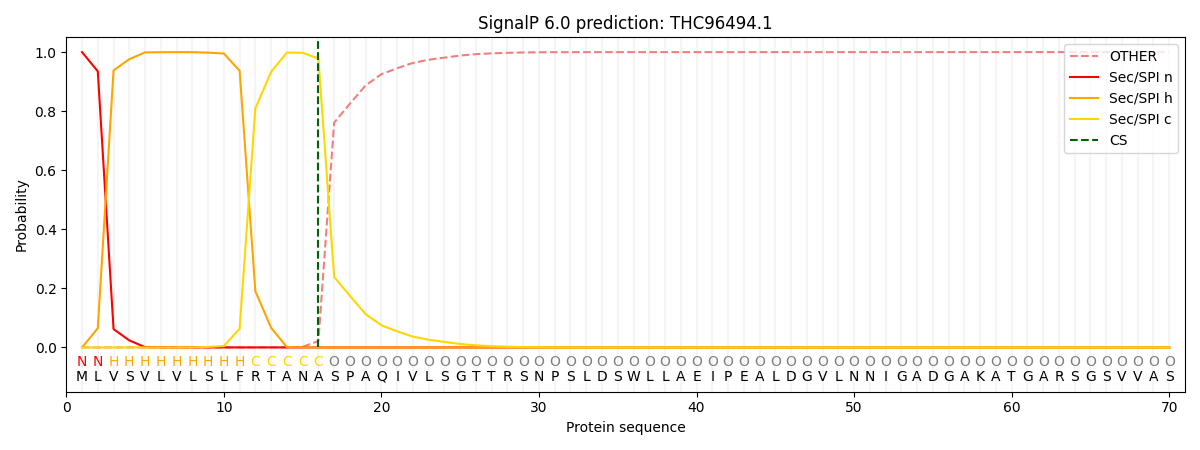
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000271 | 0.999671 | CS pos: 16-17. Pr: 0.9783 |
