You are browsing environment: FUNGIDB
CAZyme Information: THC94118.1
You are here: Home > Sequence: THC94118.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus tanneri | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus tanneri | |||||||||||
| CAZyme ID | THC94118.1 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-mannosidase [Source:UniProtKB/TrEMBL;Acc:A0A4S3JI11] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.146:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 58 | 500 | 1.7e-99 | 0.5093085106382979 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 225789 | LacZ | 6.17e-36 | 65 | 477 | 13 | 429 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| 236657 | PRK10150 | 6.15e-33 | 54 | 477 | 3 | 445 | beta-D-glucuronidase; Provisional |
| 236673 | ebgA | 4.05e-17 | 62 | 491 | 39 | 478 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| 397120 | Glyco_hydro_2_N | 6.39e-11 | 116 | 224 | 67 | 169 | Glycosyl hydrolases family 2, sugar binding domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities and has a jelly-roll fold. The domain binds the sugar moiety during the sugar-hydrolysis reaction. |
| 395572 | Glyco_hydro_2 | 4.26e-08 | 241 | 311 | 34 | 106 | Glycosyl hydrolases family 2. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 8 | 627 | 11 | 634 | |
| 0.0 | 30 | 627 | 48 | 645 | |
| 0.0 | 30 | 627 | 34 | 631 | |
| 0.0 | 30 | 627 | 34 | 631 | |
| 0.0 | 30 | 627 | 34 | 631 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.07e-107 | 38 | 615 | 12 | 577 | Chain A, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_B Chain B, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_C Chain C, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_D Chain D, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_E Chain E, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_F Chain F, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838] |
|
| 7.28e-27 | 65 | 450 | 41 | 441 | Ruminococcus gnavus Beta-glucuronidase [[Ruminococcus] gnavus],6EC6_B Ruminococcus gnavus Beta-glucuronidase [[Ruminococcus] gnavus] |
|
| 8.76e-27 | 65 | 450 | 18 | 418 | Apo structure of b-glucuronidase from Ruminococcus gnavus at 1.7 Angstrom resolution [[Ruminococcus] gnavus],6JZ1_B Apo structure of b-glucuronidase from Ruminococcus gnavus at 1.7 Angstrom resolution [[Ruminococcus] gnavus],6JZ5_A b-glucuronidase from Ruminococcus gnavus in complex with D-glucuronic acid [[Ruminococcus] gnavus],6JZ5_B b-glucuronidase from Ruminococcus gnavus in complex with D-glucuronic acid [[Ruminococcus] gnavus],6JZ6_A b-glucuronidase from Ruminococcus gnavus in complex with C6-substituted uronic isofagomine [[Ruminococcus] gnavus],6JZ6_B b-glucuronidase from Ruminococcus gnavus in complex with C6-substituted uronic isofagomine [[Ruminococcus] gnavus],6JZ7_A b-glucuronidase from Ruminococcus gnavus in complex with N1-substituted uronic isofagomine [[Ruminococcus] gnavus],6JZ7_B b-glucuronidase from Ruminococcus gnavus in complex with N1-substituted uronic isofagomine [[Ruminococcus] gnavus],6JZ8_A b-glucuronidase from Ruminococcus gnavus in complex with D-glucaro 1,5-lactone [[Ruminococcus] gnavus],6JZ8_B b-glucuronidase from Ruminococcus gnavus in complex with D-glucaro 1,5-lactone [[Ruminococcus] gnavus] |
|
| 9.74e-27 | 65 | 450 | 42 | 442 | The crystal structure of Ruminococcus gnavus beta-glucuronidase [[Ruminococcus] gnavus],5Z18_B The crystal structure of Ruminococcus gnavus beta-glucuronidase [[Ruminococcus] gnavus],5Z18_C The crystal structure of Ruminococcus gnavus beta-glucuronidase [[Ruminococcus] gnavus],5Z18_D The crystal structure of Ruminococcus gnavus beta-glucuronidase [[Ruminococcus] gnavus],5Z18_E The crystal structure of Ruminococcus gnavus beta-glucuronidase [[Ruminococcus] gnavus],5Z18_F The crystal structure of Ruminococcus gnavus beta-glucuronidase [[Ruminococcus] gnavus],5Z19_A The crystal structure of Ruminococcus gnavus beta-glucuronidase in complex with uronic isofagomine [[Ruminococcus] gnavus],5Z19_B The crystal structure of Ruminococcus gnavus beta-glucuronidase in complex with uronic isofagomine [[Ruminococcus] gnavus],5Z19_C The crystal structure of Ruminococcus gnavus beta-glucuronidase in complex with uronic isofagomine [[Ruminococcus] gnavus],5Z19_D The crystal structure of Ruminococcus gnavus beta-glucuronidase in complex with uronic isofagomine [[Ruminococcus] gnavus],5Z19_E The crystal structure of Ruminococcus gnavus beta-glucuronidase in complex with uronic isofagomine [[Ruminococcus] gnavus],5Z19_F The crystal structure of Ruminococcus gnavus beta-glucuronidase in complex with uronic isofagomine [[Ruminococcus] gnavus],6JZ2_A b-glucuronidase from Ruminococcus gnavus in complex with uronic isofagomine at 1.3 Angstroms resolution [[Ruminococcus] gnavus],6JZ2_B b-glucuronidase from Ruminococcus gnavus in complex with uronic isofagomine at 1.3 Angstroms resolution [[Ruminococcus] gnavus],6JZ3_A b-glucuronidase from Ruminococcus gnavus in complex with uronic deoxynojirimycin [[Ruminococcus] gnavus],6JZ3_B b-glucuronidase from Ruminococcus gnavus in complex with uronic deoxynojirimycin [[Ruminococcus] gnavus],6JZ4_A b-glucuronidase from Ruminococcus gnavus in complex with D-glucaro-d-lactam [[Ruminococcus] gnavus],6JZ4_B b-glucuronidase from Ruminococcus gnavus in complex with D-glucaro-d-lactam [[Ruminococcus] gnavus] |
|
| 2.30e-26 | 54 | 473 | 31 | 463 | Lactobacillus rhamnosus Beta-glucuronidase [Lacticaseibacillus rhamnosus],6ECA_B Lactobacillus rhamnosus Beta-glucuronidase [Lacticaseibacillus rhamnosus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.81e-25 | 51 | 475 | 40 | 464 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21970 PE=2 SV=1 |
|
| 1.05e-21 | 117 | 553 | 109 | 520 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22050 PE=2 SV=2 |
|
| 2.82e-19 | 116 | 445 | 58 | 382 | Beta-galactosidase OS=Thermoanaerobacter pseudethanolicus (strain ATCC 33223 / 39E) OX=340099 GN=lacZ PE=3 SV=2 |
|
| 2.91e-16 | 116 | 445 | 56 | 383 | Beta-galactosidase OS=Thermoanaerobacterium thermosulfurigenes OX=33950 GN=lacZ PE=1 SV=1 |
|
| 2.26e-15 | 65 | 477 | 13 | 441 | Beta-glucuronidase OS=Escherichia coli (strain K12) OX=83333 GN=uidA PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
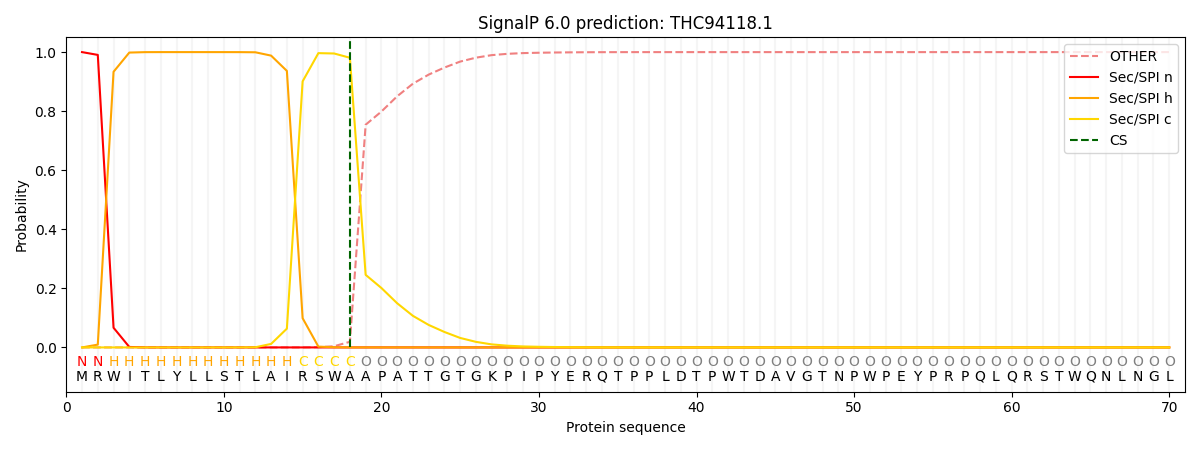
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000220 | 0.999745 | CS pos: 18-19. Pr: 0.9808 |
