You are browsing environment: FUNGIDB
CAZyme Information: THC90450.1
You are here: Home > Sequence: THC90450.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus tanneri | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus tanneri | |||||||||||
| CAZyme ID | THC90450.1 | |||||||||||
| CAZy Family | CBM87|CE18 | |||||||||||
| CAZyme Description | GMC_OxRdtase_N domain-containing protein [Source:UniProtKB/TrEMBL;Acc:A0A4S3J691] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA3 | 39 | 664 | 1.8e-91 | 0.9929577464788732 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 225186 | BetA | 3.94e-75 | 33 | 667 | 1 | 537 | Choline dehydrogenase or related flavoprotein [Lipid transport and metabolism, General function prediction only]. |
| 235000 | PRK02106 | 8.40e-74 | 36 | 667 | 2 | 535 | choline dehydrogenase; Validated |
| 398739 | GMC_oxred_C | 3.77e-32 | 512 | 659 | 2 | 143 | GMC oxidoreductase. This domain found associated with pfam00732. |
| 215420 | PLN02785 | 1.06e-19 | 15 | 645 | 32 | 558 | Protein HOTHEAD |
| 366272 | GMC_oxred_N | 1.91e-15 | 140 | 413 | 19 | 218 | GMC oxidoreductase. This family of proteins bind FAD as a cofactor. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 667 | 1 | 677 | |
| 0.0 | 1 | 667 | 1 | 677 | |
| 0.0 | 1 | 667 | 1 | 677 | |
| 0.0 | 1 | 667 | 1 | 658 | |
| 1.63e-315 | 51 | 667 | 52 | 664 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.46e-39 | 39 | 663 | 13 | 526 | Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_B Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_C Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_D Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_E Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_F Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_G Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_H Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis] |
|
| 7.46e-39 | 39 | 663 | 13 | 526 | Chain A, Choline oxidase [Arthrobacter globiformis],3LJP_B Chain B, Choline oxidase [Arthrobacter globiformis] |
|
| 1.86e-38 | 39 | 663 | 13 | 526 | Crystal structure of choline oxidase reveals insights into the catalytic mechanism [Arthrobacter globiformis],2JBV_B Crystal structure of choline oxidase reveals insights into the catalytic mechanism [Arthrobacter globiformis],4MJW_A Crystal Structure of Choline Oxidase in Complex with the Reaction Product Glycine Betaine [Arthrobacter globiformis],4MJW_B Crystal Structure of Choline Oxidase in Complex with the Reaction Product Glycine Betaine [Arthrobacter globiformis] |
|
| 3.04e-32 | 39 | 666 | 1 | 564 | Crystal structure of aryl-alcohol oxidase from Pleurotus eryngii in complex with p-anisic acid [Pleurotus eryngii] |
|
| 4.12e-32 | 38 | 666 | 1 | 565 | Crystal structure of aryl-alcohol-oxidase from Pleurotus eryingii [Pleurotus eryngii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.68e-44 | 39 | 665 | 2 | 527 | Oxygen-dependent choline dehydrogenase OS=Brucella anthropi (strain ATCC 49188 / DSM 6882 / CCUG 24695 / JCM 21032 / LMG 3331 / NBRC 15819 / NCTC 12168 / Alc 37) OX=439375 GN=betA PE=3 SV=1 |
|
| 1.19e-43 | 38 | 663 | 39 | 607 | Dehydrogenase pkfF OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=pkfF PE=2 SV=1 |
|
| 1.79e-43 | 40 | 665 | 8 | 540 | Oxygen-dependent choline dehydrogenase OS=Staphylococcus epidermidis (strain ATCC 35984 / RP62A) OX=176279 GN=betA PE=3 SV=1 |
|
| 1.79e-43 | 40 | 665 | 8 | 540 | Oxygen-dependent choline dehydrogenase OS=Staphylococcus epidermidis (strain ATCC 12228 / FDA PCI 1200) OX=176280 GN=betA PE=3 SV=1 |
|
| 2.34e-43 | 35 | 665 | 3 | 538 | Oxygen-dependent choline dehydrogenase OS=Staphylococcus aureus (strain MW2) OX=196620 GN=betA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
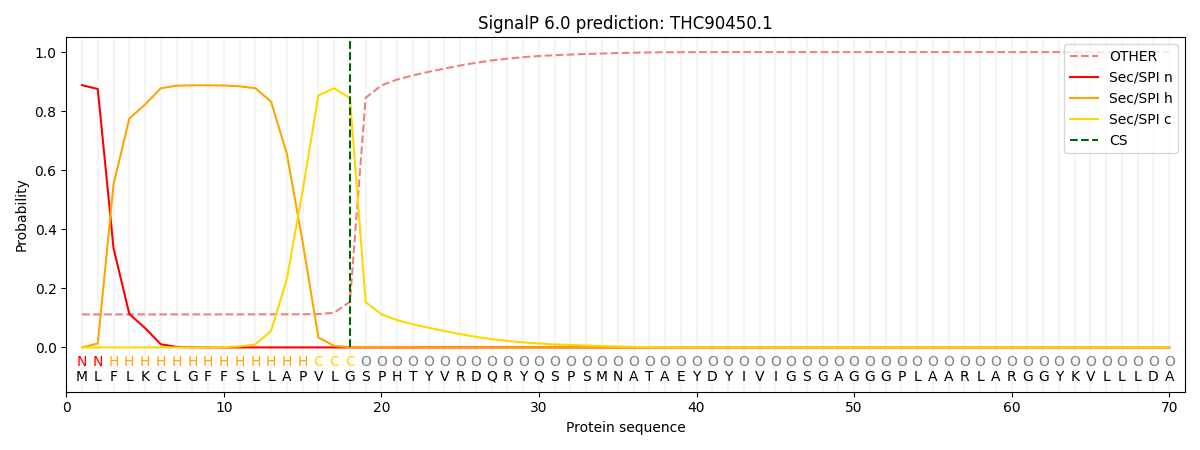
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.118500 | 0.881474 | CS pos: 18-19. Pr: 0.8447 |
