You are browsing environment: FUNGIDB
CAZyme Information: SSD61017.1
You are here: Home > Sequence: SSD61017.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Saccharomycodes ludwigii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Saccharomycetes; ; Saccharomycodaceae; Saccharomycodes; Saccharomycodes ludwigii | |||||||||||
| CAZyme ID | SSD61017.1 | |||||||||||
| CAZy Family | GT22 | |||||||||||
| CAZyme Description | Glyco_hydro_15 domain-containing protein [Source:UniProtKB/TrEMBL;Acc:A0A376B8N3] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH15 | 169 | 659 | 1.1e-74 | 0.961218836565097 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395586 | Glyco_hydro_15 | 6.43e-66 | 156 | 653 | 5 | 406 | Glycosyl hydrolases family 15. In higher organisms this family is represented by phosphorylase kinase subunits. |
| 225922 | SGA1 | 8.60e-15 | 252 | 653 | 245 | 591 | Glucoamylase (glucan-1,4-alpha-glucosidase), GH15 family [Carbohydrate transport and metabolism]. |
| 273702 | oligosac_amyl | 5.15e-05 | 356 | 565 | 388 | 556 | oligosaccharide amylase. The name of this type of amylase is based on the characterization of an glucoamylase family enzyme from Thermoactinomyces vulgaris. The T. vulgaris enzyme was expressed in E. coli and, like other glucoamylases, it releases beta-D-glucose from starch. However, unlike previously characterized glucoamylases, this T. vulgaris amylase hydrolyzes maltooligosaccharides (maltotetraose, maltose) more efficiently than starch (1), indicating this enzyme belongs to a class of glucoamylase-type enzymes with oligosaccharide-metabolizing activity. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 7.76e-187 | 103 | 674 | 64 | 609 | |
| 1.92e-176 | 140 | 662 | 100 | 610 | |
| 4.23e-175 | 125 | 667 | 86 | 594 | |
| 1.47e-172 | 140 | 674 | 118 | 650 | |
| 4.00e-172 | 141 | 670 | 97 | 601 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.68e-91 | 137 | 655 | 12 | 474 | Chain A, GLUCOAMYLASE [Saccharomycopsis fibuligera],2F6D_A Chain A, Glucoamylase GLU1 [Saccharomycopsis fibuligera],2FBA_A Chain A, Glucoamylase GLU1 [Saccharomycopsis fibuligera] |
|
| 7.97e-54 | 136 | 659 | 24 | 454 | Chain A, Glucoamylase P [Amorphotheca resinae],6FHW_B Chain B, Glucoamylase P [Amorphotheca resinae] |
|
| 1.37e-48 | 146 | 666 | 5 | 433 | Chain A, GLUCOAMYLASE [Trichoderma reesei],2VN7_A Chain A, GLUCOAMYLASE [Trichoderma reesei] |
|
| 7.67e-46 | 141 | 658 | 1 | 420 | Refined structure for the complex of acarbose with glucoamylase from Aspergillus awamori var. x100 to 2.4 angstroms resolution [Aspergillus awamori],1DOG_A REFINED STRUCTURE FOR THE COMPLEX OF 1-DEOXYNOJIRIMYCIN WITH GLUCOAMYLASE FROM (ASPERGILLUS AWAMORI) VAR. X100 TO 2.4 ANGSTROMS RESOLUTION [Aspergillus awamori],1GLM_A REFINED CRYSTAL STRUCTURES OF GLUCOAMYLASE FROM ASPERGILLUS AWAMORI VAR. X100 [Aspergillus awamori],3GLY_A REFINED CRYSTAL STRUCTURES OF GLUCOAMYLASE FROM ASPERGILLUS AWAMORI VAR. X100 [Aspergillus awamori] |
|
| 7.82e-46 | 141 | 658 | 1 | 420 | GLUCOAMYLASE-471 COMPLEXED WITH ACARBOSE [Aspergillus awamori] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.79e-117 | 146 | 662 | 84 | 538 | Glucoamylase, intracellular sporulation-specific OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=SGA1 PE=3 SV=2 |
|
| 2.75e-96 | 146 | 634 | 342 | 768 | Glucoamylase S2 OS=Saccharomyces cerevisiae OX=4932 GN=STA2 PE=3 SV=1 |
|
| 1.43e-95 | 146 | 634 | 341 | 767 | Glucoamylase S1 OS=Saccharomyces cerevisiae OX=4932 GN=STA1 PE=3 SV=2 |
|
| 9.46e-90 | 137 | 655 | 39 | 501 | Glucoamylase GLU1 OS=Saccharomycopsis fibuligera OX=4944 GN=GLU1 PE=1 SV=1 |
|
| 1.98e-88 | 137 | 655 | 39 | 501 | Glucoamylase GLA1 OS=Saccharomycopsis fibuligera OX=4944 GN=GLA1 PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
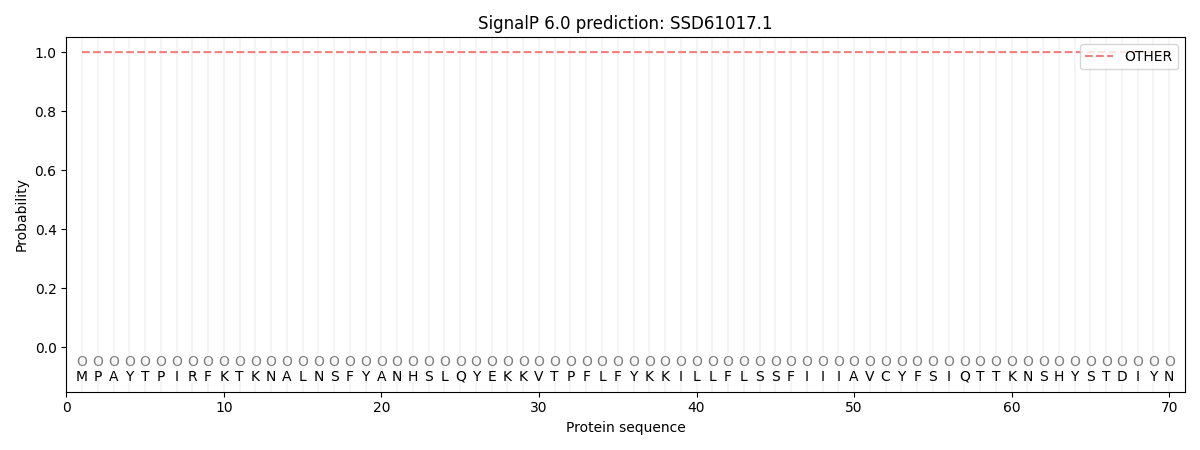
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999980 | 0.000022 |

