You are browsing environment: FUNGIDB
CAZyme Information: SPSK_09288-t39_1-p1
You are here: Home > Sequence: SPSK_09288-t39_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Sporothrix schenckii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Ophiostomataceae; Sporothrix; Sporothrix schenckii | |||||||||||
| CAZyme ID | SPSK_09288-t39_1-p1 | |||||||||||
| CAZy Family | GT25 | |||||||||||
| CAZyme Description | oxidoreductase domain-containing protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.49:6 | 3.2.1.52:2 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH109 | 9 | 172 | 2.4e-26 | 0.40852130325814534 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 223745 | MviM | 1.37e-40 | 6 | 235 | 2 | 224 | Predicted dehydrogenase [General function prediction only]. |
| 397161 | GFO_IDH_MocA_C | 7.02e-18 | 148 | 435 | 1 | 204 | Oxidoreductase family, C-terminal alpha/beta domain. This family of enzymes utilize NADP or NAD. This family is called the GFO/IDH/MOCA family in swiss-prot. |
| 396129 | GFO_IDH_MocA | 1.55e-13 | 8 | 136 | 1 | 120 | Oxidoreductase family, NAD-binding Rossmann fold. This family of enzymes utilize NADP or NAD. This family is called the GFO/IDH/MOCA family in swiss-prot. |
| 182305 | PRK10206 | 1.72e-04 | 60 | 166 | 42 | 154 | putative oxidoreductase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.12e-25 | 9 | 392 | 396 | 709 | |
| 2.59e-10 | 3 | 159 | 61 | 219 | |
| 5.68e-09 | 7 | 159 | 54 | 206 | |
| 5.95e-09 | 9 | 193 | 76 | 267 | |
| 5.95e-09 | 9 | 193 | 76 | 267 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.26e-10 | 65 | 216 | 60 | 208 | Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with glycerol [Caulobacter vibrioides CB15],5A02_B Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with glycerol [Caulobacter vibrioides CB15],5A02_C Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with glycerol [Caulobacter vibrioides CB15],5A02_D Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with glycerol [Caulobacter vibrioides CB15],5A02_E Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with glycerol [Caulobacter vibrioides CB15],5A02_F Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with glycerol [Caulobacter vibrioides CB15],5A03_A Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with xylose [Caulobacter vibrioides CB15],5A03_B Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with xylose [Caulobacter vibrioides CB15],5A03_C Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with xylose [Caulobacter vibrioides CB15],5A03_D Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with xylose [Caulobacter vibrioides CB15],5A03_E Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with xylose [Caulobacter vibrioides CB15],5A03_F Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with xylose [Caulobacter vibrioides CB15],5A04_A Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with glucose [Caulobacter vibrioides CB15],5A04_B Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with glucose [Caulobacter vibrioides CB15],5A04_C Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with glucose [Caulobacter vibrioides CB15],5A04_D Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with glucose [Caulobacter vibrioides CB15],5A04_E Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with glucose [Caulobacter vibrioides CB15],5A04_F Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with glucose [Caulobacter vibrioides CB15],5A05_A Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with maltotriose [Caulobacter vibrioides CB15],5A05_B Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with maltotriose [Caulobacter vibrioides CB15],5A05_C Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with maltotriose [Caulobacter vibrioides CB15],5A05_D Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with maltotriose [Caulobacter vibrioides CB15],5A05_E Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with maltotriose [Caulobacter vibrioides CB15],5A05_F Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with maltotriose [Caulobacter vibrioides CB15],5A06_A Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with sorbitol [Caulobacter vibrioides CB15],5A06_B Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with sorbitol [Caulobacter vibrioides CB15],5A06_C Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with sorbitol [Caulobacter vibrioides CB15],5A06_D Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with sorbitol [Caulobacter vibrioides CB15],5A06_E Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with sorbitol [Caulobacter vibrioides CB15],5A06_F Crystal structure of aldose-aldose oxidoreductase from Caulobacter crescentus complexed with sorbitol [Caulobacter vibrioides CB15] |
|
| 7.87e-10 | 68 | 213 | 55 | 196 | CRYSTAL STRUCTURE OF NAD-BINDING PROTEIN FROM Listeria innocua [Listeria innocua],3E18_B CRYSTAL STRUCTURE OF NAD-BINDING PROTEIN FROM Listeria innocua [Listeria innocua] |
|
| 2.07e-09 | 9 | 218 | 4 | 202 | The crystal structure of the putative dehydrogenase from Bordetella bronchiseptica RB50 [Bordetella bronchiseptica] |
|
| 2.41e-09 | 68 | 222 | 67 | 220 | Chain A, Inositol dehydrogenase [Lacticaseibacillus casei BL23],4N54_B Chain B, Inositol dehydrogenase [Lacticaseibacillus casei BL23],4N54_C Chain C, Inositol dehydrogenase [Lacticaseibacillus casei BL23],4N54_D Chain D, Inositol dehydrogenase [Lacticaseibacillus casei BL23] |
|
| 2.44e-09 | 68 | 222 | 70 | 223 | Chain A, Inositol dehydrogenase [Lacticaseibacillus casei BL23],4MKZ_A Chain A, Inositol dehydrogenase [Lacticaseibacillus casei BL23] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.59e-75 | 7 | 438 | 1 | 422 | Putative oxidoreductase YteT OS=Bacillus subtilis (strain 168) OX=224308 GN=yteT PE=2 SV=1 |
|
| 1.32e-11 | 9 | 275 | 3 | 250 | Uncharacterized oxidoreductase SP_1686 OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=SP_1686 PE=3 SV=2 |
|
| 3.43e-10 | 78 | 213 | 62 | 190 | Inositol 2-dehydrogenase OS=Rhizobium meliloti (strain 1021) OX=266834 GN=idhA PE=1 SV=2 |
|
| 1.40e-09 | 9 | 193 | 76 | 267 | Glycosyl hydrolase family 109 protein 1 OS=Akkermansia muciniphila (strain ATCC BAA-835 / DSM 22959 / JCM 33894 / BCRC 81048 / CCUG 64013 / CIP 107961 / Muc) OX=349741 GN=Amuc_0017 PE=3 SV=1 |
|
| 3.13e-09 | 7 | 159 | 54 | 206 | Glycosyl hydrolase family 109 protein OS=Shewanella baltica (strain OS185) OX=402882 GN=Shew185_2813 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
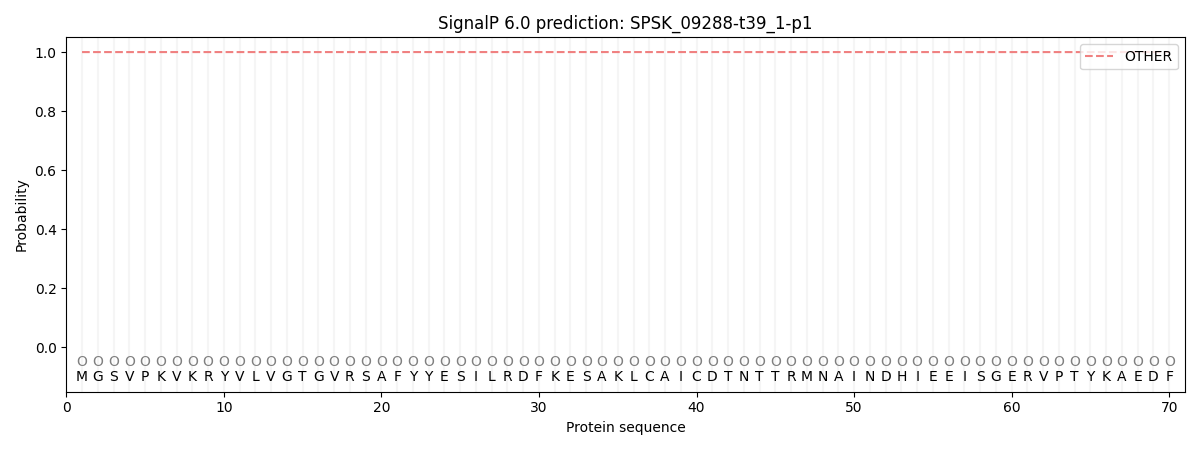
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000092 | 0.000000 |
