You are browsing environment: FUNGIDB
CAZyme Information: SMR47185.1
You are here: Home > Sequence: SMR47185.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Zymoseptoria tritici | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Mycosphaerellaceae; Zymoseptoria; Zymoseptoria tritici | |||||||||||
| CAZyme ID | SMR47185.1 | |||||||||||
| CAZy Family | CE4 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.25:3 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 6 | 731 | 7.1e-85 | 0.6954787234042553 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 225789 | LacZ | 1.48e-72 | 3 | 660 | 7 | 639 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| 395572 | Glyco_hydro_2 | 2.24e-13 | 194 | 298 | 1 | 106 | Glycosyl hydrolases family 2. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
| 407658 | Mannosidase_ig | 6.80e-10 | 676 | 763 | 2 | 91 | Mannosidase Ig/CBM-like domain. This domain corresponds to domain 4 in the structure of Bacteroides thetaiotaomicron beta-mannosidase, BtMan2A. This domain has an Ig-like fold. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 836 | 1 | 836 | |
| 0.0 | 1 | 836 | 1 | 836 | |
| 0.0 | 1 | 836 | 1 | 836 | |
| 0.0 | 5 | 836 | 1 | 832 | |
| 0.0 | 1 | 836 | 10 | 845 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.87e-140 | 5 | 658 | 24 | 661 | Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12],5N6U_B Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12],5N6U_C Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12],5N6U_D Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12] |
|
| 6.89e-120 | 5 | 818 | 3 | 832 | Crystal structure of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri in complex with mannose [Xanthomonas citri pv. citri str. 306],6BYE_B Crystal structure of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri in complex with mannose [Xanthomonas citri pv. citri str. 306] |
|
| 7.05e-120 | 5 | 818 | 3 | 832 | Crystal structure of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306] |
|
| 4.92e-119 | 5 | 818 | 1 | 830 | Crystal structure of the acid-base mutant (E477A) of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306],6BYI_B Crystal structure of the acid-base mutant (E477A) of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306] |
|
| 5.04e-119 | 5 | 818 | 3 | 832 | Crystal structure of the nucleophile mutant (E575A) of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306],6BYG_B Crystal structure of the nucleophile mutant (E575A) of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 10 | 835 | 9 | 844 | Beta-mannosidase B OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=mndB PE=3 SV=3 |
|
| 0.0 | 10 | 835 | 9 | 843 | Beta-mannosidase B OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=mndB PE=1 SV=2 |
|
| 0.0 | 9 | 835 | 8 | 843 | Beta-mannosidase B OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=mndB PE=3 SV=2 |
|
| 0.0 | 9 | 835 | 8 | 844 | Beta-mannosidase B OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=mndB PE=3 SV=1 |
|
| 0.0 | 5 | 835 | 4 | 845 | Beta-mannosidase B OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=mndB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
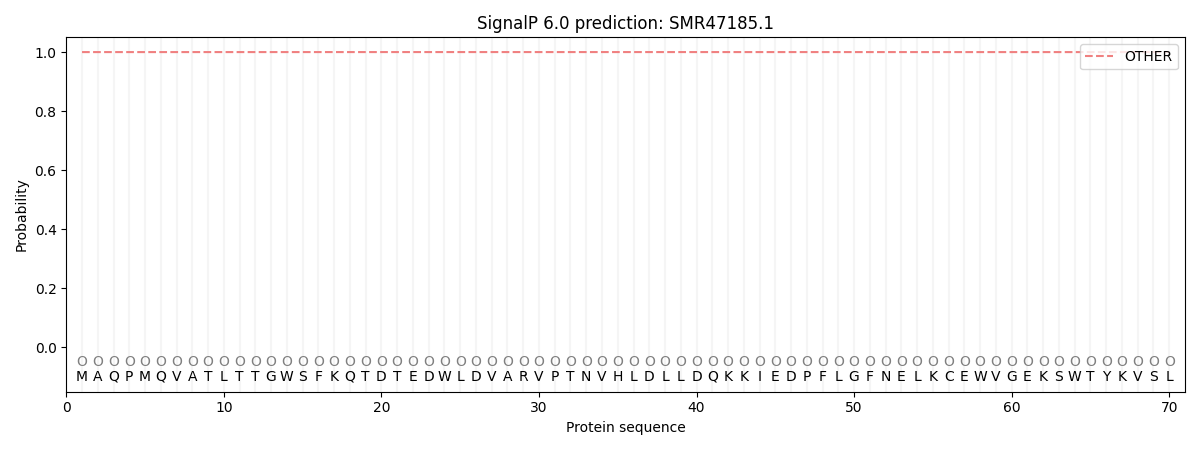
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000075 | 0.000000 |
