You are browsing environment: FUNGIDB
CAZyme Information: SMR45125.1
You are here: Home > Sequence: SMR45125.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Zymoseptoria tritici | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Mycosphaerellaceae; Zymoseptoria; Zymoseptoria tritici | |||||||||||
| CAZyme ID | SMR45125.1 | |||||||||||
| CAZy Family | AA9 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH64 | 38 | 409 | 1.3e-93 | 0.989100817438692 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 185759 | GH64-GluB-like | 2.33e-162 | 39 | 410 | 1 | 369 | glycoside hydrolase family 64: beta-1,3-glucanase B (GluB)-like. This subfamily is represented by GluB, beta-1,3-glucanase B , from Lysobacter enzymogenes Strain N4-7 and related bacterial and ascomycete proteins. GluB is a member of the glycoside hydrolase family 64 (GH64) involved in the cleavage of long-chain polysaccharide beta-1,3-glucans, into specific pentasaccharide oligomers. Among bacteria, many beta-1,3-glucanases are implicated in fungal cell wall degradation. GluB possesses the conserved Glu and Asp residues required to cleave substrate beta-1,3-glucans. Recombinant GluB demonstrated higher relative activity toward the branched-chain beta-1,3 glucan substrate zymosan A than toward linear beta-1,3 glucan substrates. Based on the structure of laminaripentaose-producing, beta-1,3-glucanase (LPHase) of Streptomyces matensis, which belongs to the same family as GluB but to a different subfamily, this cd is a two-domain model. Sometimes these two domains are found associated with other domains such as in the Catenulispora acidiphila DSM 44928 carbohydrate binding family 6 protein in which they are positioned N-terminal of a carbohydrate binding module, family 6 (CBM_6) domain. |
| 406796 | Glyco_hydro_64 | 2.18e-161 | 39 | 409 | 3 | 370 | Beta-1,3-glucanase. Family 64 glycoside hydrolases have beta-1,3-glucanase activity. |
| 185755 | GH64-LPHase-like | 4.17e-72 | 39 | 407 | 1 | 350 | glycoside hydrolase family 64: laminaripentaose-producing, beta-1,3-glucanase (LPHase)-like. This subfamily is represented by the laminaripentaose-producing, beta-1,3-glucanase (LPHase) of Streptomyces matensis and related bacterial and ascomycete proteins. LPHase is a member of glycoside hydrolase family 64 (GH64), it is an inverting enzyme involved in the cleavage of long-chain polysaccharide beta-1,3-glucans, into specific pentasaccharide oligomers. LPHase is a two-domain crescent fold structure: one domain is composed of 10 beta-strands, eight coming from the N-terminus of the protein and two from the C-terminal region, and the protein has a second inserted domain; this cd includes both domains. This protein has an electronegative, substrate-binding cleft, and conserved Glu and Asp residues involved in the cleavage of the beta-1,3-glucan, laminarin, a plant and fungal cell wall component. Among bacteria, many beta-1,3-glucanases are implicated in fungal cell wall degradation. Also included in this family is GluB , the beta-1,3-glucanase B from Lysobacter enzymogenes Strain N4-7. Recombinant GluB demonstrated higher relative activity toward the branched-chain beta-1,3 glucan substrate zymosan A than toward linear beta-1,3 glucan substrates. Sometimes these two domains are found associated with other domains such as in the Catenulispora acidiphila DSM 44928 carbohydrate binding family 6 protein in which they are positioned N-terminal of a carbohydrate binding module, family 6 (CBM_6) domain. In the Cellulosimicrobium cellulans, glucan endo-1,3-beta-glucosidase, they are positioned N-terminal of a RICIN, carbohydrate-binding domain. |
| 185753 | GH64-like | 3.29e-41 | 39 | 410 | 1 | 319 | glycosyl hydrolase 64 family. This family is represented by the laminaripentaose-producing, beta-1,3-glucanase (LPHase) of Streptomyces matensis and related bacterial and ascomycete proteins. LPHase is a member of glycoside hydrolase family 64 (GH64), it is an inverting enzyme involved in the cleavage of long-chain polysaccharide beta-1,3-glucans, into specific pentasaccharide oligomers. LPHase is a two-domain crescent fold structure: one domain is composed of 10 beta-strands, eight coming from the N-terminus of the protein and two from the C-terminal region, and the protein has a second inserted domain; this cd includes both domains. This protein has an electronegative, substrate-binding cleft, and conserved Glu and Asp residues involved in the cleavage of the beta-1,3-glucan, laminarin, a plant and fungal cell wall component. Among bacteria, many beta-1,3-glucanases are implicated in fungal cell wall degradation. Also included in this family is GluB , the beta-1,3-glucanase B from Lysobacter enzymogenes Strain N4-7. Recombinant GluB demonstrated higher relative activity toward the branched-chain beta-1,3 glucan substrate zymosan A than toward linear beta-1,3 glucan substrates. Sometimes these two domains are found associated with other domains such as in the Catenulispora acidiphila DSM 44928 carbohydrate binding family 6 protein in which they are positioned N-terminal of a carbohydrate binding module, family 6 (CBM_6) domain. In the Cellulosimicrobium cellulans, glucan endo-1,3-beta-glucosidase, they are positioned N-terminal of a RICIN, carbohydrate-binding domain, and in the Salinispora tropica CNB-440, coagulation factor 5/8 C-terminal domain (FA58C) protein, they are positioned C-terminal of two FA58C domains which are proposed to function as cell surface-attached, carbohydrate-binding domain. This FA58C-containing protein has an internal peptide deletion (of approx. 44 residues) in the LPHase domain II. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 6.82e-313 | 1 | 412 | 1 | 412 | |
| 3.95e-312 | 1 | 412 | 1 | 412 | |
| 5.43e-311 | 1 | 412 | 33 | 444 | |
| 6.33e-310 | 1 | 412 | 33 | 444 | |
| 7.86e-219 | 2 | 411 | 36 | 454 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
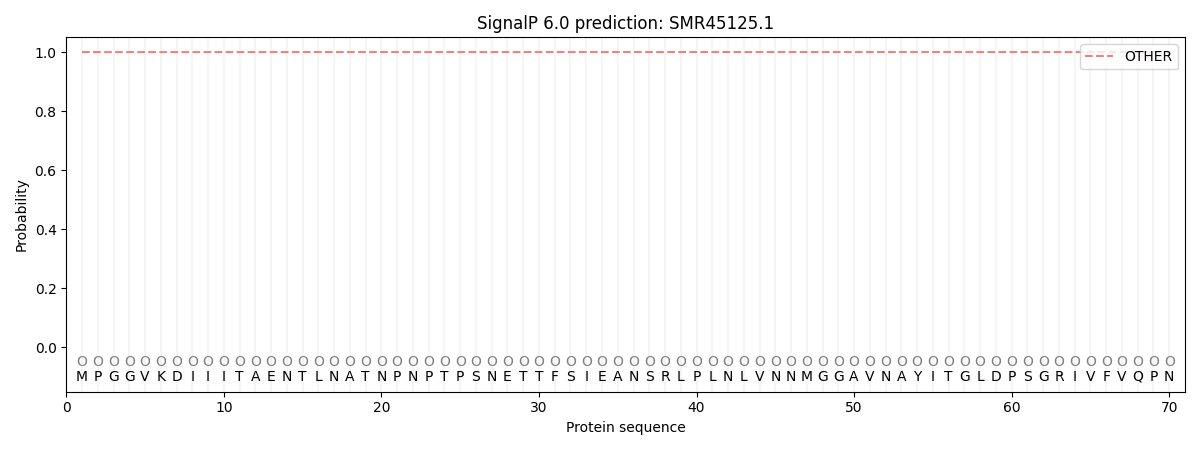
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999974 | 0.000063 |
