You are browsing environment: FUNGIDB
CAZyme Information: SDRG_04820-t26_1-p1
You are here: Home > Sequence: SDRG_04820-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Saprolegnia diclina | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Saprolegniaceae; Saprolegnia; Saprolegnia diclina | |||||||||||
| CAZyme ID | SDRG_04820-t26_1-p1 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 53 | 327 | 5.7e-49 | 0.7325581395348837 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 341270 | ACS | 0.010 | 372 | 410 | 115 | 145 | Acetyl-CoA synthetase (also known as acetate-CoA ligase and acetyl-activating enzyme). Acetyl-CoA synthetase (ACS, EC 6.2.1.1, acetate#CoA ligase or acetate:CoA ligase (AMP-forming)) catalyzes the formation of acetyl-CoA from acetate, CoA, and ATP. Synthesis of acetyl-CoA is carried out in a two-step reaction. In the first step, the enzyme catalyzes the synthesis of acetyl-AMP intermediate from acetate and ATP. In the second step, acetyl-AMP reacts with CoA to produce acetyl-CoA. This enzyme is widely present in all living organisms. The activity of this enzyme is crucial for maintaining the required levels of acetyl-CoA, a key intermediate in many important biosynthetic and catabolic processes. Acetyl-CoA is used in the biosynthesis of glucose, fatty acids, and cholesterol. It can also be used in the production of energy in the citric acid cycle. Eukaryotes typically have two isoforms of acetyl-CoA synthetase, a cytosolic form involved in biosynthetic processes and a mitochondrial form primarily involved in energy generation. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.24e-38 | 27 | 499 | 140 | 567 | |
| 1.19e-34 | 23 | 496 | 139 | 567 | |
| 6.43e-33 | 23 | 496 | 138 | 566 | |
| 2.76e-32 | 29 | 496 | 162 | 578 | |
| 7.52e-28 | 23 | 496 | 135 | 563 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.67e-11 | 34 | 334 | 41 | 320 | Crystal analysis of the complex structure, E201A-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii],3QHN_B Crystal analysis of the complex structure, E201A-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii],3QHN_C Crystal analysis of the complex structure, E201A-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii] |
|
| 5.09e-11 | 34 | 334 | 8 | 287 | The Crystal Structure of Cellulase-Inhibitor Complex. [Pyrococcus horikoshii OT3],3VVG_B The Crystal Structure of Cellulase-Inhibitor Complex. [Pyrococcus horikoshii OT3],3VVG_C The Crystal Structure of Cellulase-Inhibitor Complex. [Pyrococcus horikoshii OT3],3W6L_A Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],3W6L_B Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],3W6L_C Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM1_A Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM1_B Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM1_C Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3] |
|
| 6.83e-11 | 34 | 334 | 41 | 320 | Functional Analysis of Hyperthermophilic Endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3],2ZUN_A Functional Analysis of Hyperthermophilic Endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3],2ZUN_B Functional Analysis of Hyperthermophilic Endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3],2ZUN_C Functional Analysis of Hyperthermophilic Endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3] |
|
| 6.83e-11 | 34 | 334 | 41 | 320 | Functional analysis of hyperthermophilic endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3],3AXX_B Functional analysis of hyperthermophilic endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3],3AXX_C Functional analysis of hyperthermophilic endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3] |
|
| 6.83e-11 | 34 | 334 | 41 | 320 | Crystal analysis of the complex structure, E342A-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii],3QHM_B Crystal analysis of the complex structure, E342A-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii],3QHM_C Crystal analysis of the complex structure, E342A-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
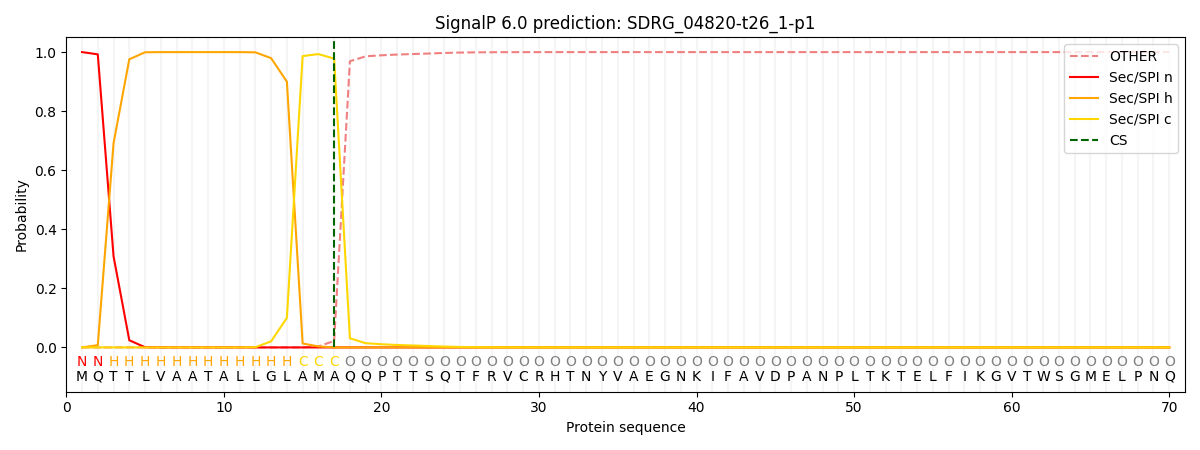
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000219 | 0.999766 | CS pos: 17-18. Pr: 0.9780 |
