You are browsing environment: FUNGIDB
CAZyme Information: SAPIO_CDS6484-t41_1-p1
You are here: Home > Sequence: SAPIO_CDS6484-t41_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Scedosporium apiospermum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Microascaceae; Scedosporium; Scedosporium apiospermum | |||||||||||
| CAZyme ID | SAPIO_CDS6484-t41_1-p1 | |||||||||||
| CAZy Family | GT71 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.106:25 | 3.2.1.-:4 |
|---|
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397353 | Glyco_hydro_63 | 5.17e-90 | 290 | 605 | 1 | 364 | Glycosyl hydrolase family 63 C-terminal domain. This is a family of eukaryotic enzymes belonging to glycosyl hydrolase family 63. They catalyze the specific cleavage of the non-reducing terminal glucose residue from Glc(3)Man(9)GlcNAc(2). Mannosyl oligosaccharide glucosidase EC:3.2.1.106 is the first enzyme in the N-linked oligosaccharide processing pathway. This family represents the C-terminal catalytic domain. |
| 407154 | Glyco_hydro_63N | 2.13e-51 | 87 | 202 | 3 | 120 | Glycosyl hydrolase family 63 N-terminal domain. This is a family of eukaryotic enzymes belonging to glycosyl hydrolase family 63. They catalyze the specific cleavage of the non-reducing terminal glucose residue from Glc(3)Man(9)GlcNAc(2). Mannosyl oligosaccharide glucosidase EC:3.2.1.106 is the first enzyme in the N-linked oligosaccharide processing pathway. This family represents the N-terminal beta sandwich domain. |
| 293786 | ANK | 2.99e-04 | 702 | 750 | 48 | 96 | ankyrin repeats. Ankyrin repeats are one of the most abundant repeat motifs, and generally function as scaffolds for protein-protein interactions in processes including cell cycle, transcriptional regulation, signal transduction, vesicular trafficking, and inflammatory response. Although predominantly found in eukaryotic proteins, they are also found in some bacterial and viral proteins. Less is known of their physiological roles in prokaryotes. Some bacterial ANK proteins play key roles in microbial pathogenesis by mimicking or manipulating host function(s). The pathogen Providencia alcalifaciens N-formyltransferase ankyrin repeats function in small molecule binding and allosteric control. Ankyrin-repeat proteins have been associated with a number of human diseases. |
| 372654 | Ank_4 | 4.16e-04 | 702 | 741 | 14 | 53 | Ankyrin repeats (many copies). |
| 293786 | ANK | 4.96e-04 | 702 | 762 | 15 | 71 | ankyrin repeats. Ankyrin repeats are one of the most abundant repeat motifs, and generally function as scaffolds for protein-protein interactions in processes including cell cycle, transcriptional regulation, signal transduction, vesicular trafficking, and inflammatory response. Although predominantly found in eukaryotic proteins, they are also found in some bacterial and viral proteins. Less is known of their physiological roles in prokaryotes. Some bacterial ANK proteins play key roles in microbial pathogenesis by mimicking or manipulating host function(s). The pathogen Providencia alcalifaciens N-formyltransferase ankyrin repeats function in small molecule binding and allosteric control. Ankyrin-repeat proteins have been associated with a number of human diseases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.88e-223 | 73 | 618 | 31 | 674 | |
| 1.99e-213 | 84 | 618 | 42 | 656 | |
| 7.17e-185 | 55 | 618 | 1 | 650 | |
| 6.15e-147 | 87 | 619 | 36 | 667 | |
| 4.46e-143 | 87 | 618 | 39 | 683 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.91e-74 | 79 | 619 | 31 | 661 | Murine endoplasmic reticulum alpha-glucosidase I with N-9'-methoxynonyl-1-deoxynojirimycin [Mus musculus],5MHF_B Murine endoplasmic reticulum alpha-glucosidase I with N-9'-methoxynonyl-1-deoxynojirimycin [Mus musculus],5MHF_C Murine endoplasmic reticulum alpha-glucosidase I with N-9'-methoxynonyl-1-deoxynojirimycin [Mus musculus],5MHF_D Murine endoplasmic reticulum alpha-glucosidase I with N-9'-methoxynonyl-1-deoxynojirimycin [Mus musculus] |
|
| 4.77e-67 | 87 | 618 | 17 | 686 | Crystal structure of Processing alpha-Glucosidase I [Saccharomyces cerevisiae S288C] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.57e-96 | 87 | 616 | 44 | 691 | Probable mannosyl-oligosaccharide glucosidase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPAC6G10.09 PE=3 SV=1 |
|
| 1.65e-74 | 79 | 619 | 86 | 716 | Mannosyl-oligosaccharide glucosidase OS=Mus musculus OX=10090 GN=Mogs PE=1 SV=1 |
|
| 6.40e-70 | 79 | 619 | 86 | 716 | Mannosyl-oligosaccharide glucosidase OS=Rattus norvegicus OX=10116 GN=Mogs PE=1 SV=1 |
|
| 2.78e-67 | 79 | 619 | 88 | 719 | Mannosyl-oligosaccharide glucosidase OS=Homo sapiens OX=9606 GN=MOGS PE=1 SV=5 |
|
| 3.31e-66 | 87 | 618 | 47 | 716 | Mannosyl-oligosaccharide glucosidase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=CWH41 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
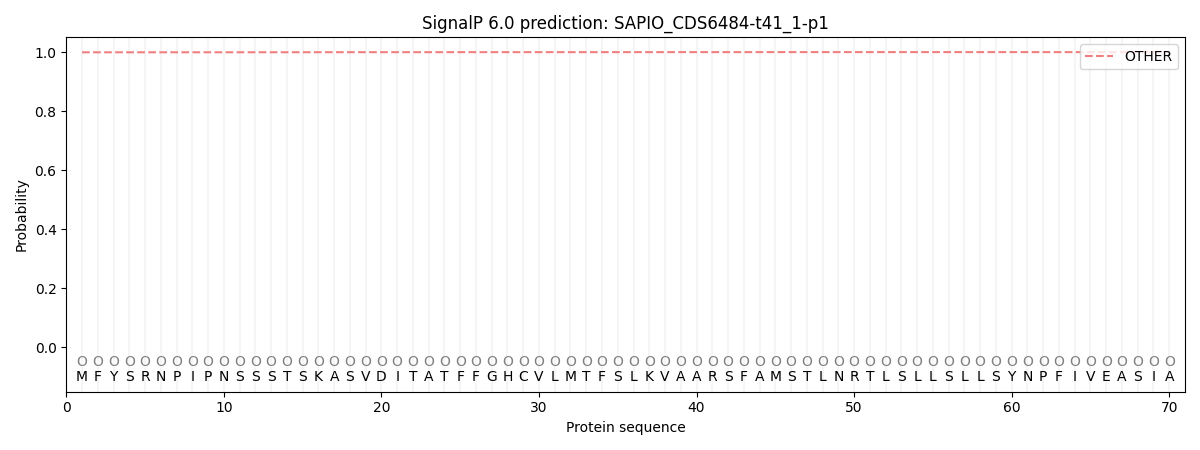
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999413 | 0.000592 |

