You are browsing environment: FUNGIDB
CAZyme Information: SAPIO_CDS5444-t41_1-p1
You are here: Home > Sequence: SAPIO_CDS5444-t41_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Scedosporium apiospermum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Microascaceae; Scedosporium; Scedosporium apiospermum | |||||||||||
| CAZyme ID | SAPIO_CDS5444-t41_1-p1 | |||||||||||
| CAZy Family | GH20 | |||||||||||
| CAZyme Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.113:12 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 64 | 538 | 4.4e-126 | 0.9955156950672646 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 2.34e-180 | 63 | 538 | 1 | 453 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 2.22e-113 | 53 | 539 | 69 | 519 | glycoside hydrolase family 47 protein; Provisional |
| 319897 | Col_Im_like | 0.004 | 154 | 225 | 14 | 71 | inhibitory immunity (Im) protein of colicin (Col) deoxyribonuclease (DNase) and pyocins. This family contains inhibitory immunity (Im) proteins that bind to colicin endonucleases (DNases) or pyocins with very high affinity and specificity; this is critical for the neutralization of endogenous DNase catalytic activity and for protection against exogenous DNase bacteriocins. The DNase colicin family (ColE2, ColE7, ColE8 and ColE9) in E. coli, and pyocin family (S1, S2, S3 and AP41) in P. aeruginosa, are potent bacteriocins where the immunity proteins (Ims) protect the colicin/pyocin producing (i.e. colicinogenic) bacteria by binding and inactivating colicin nucleases. The binding affinities between cognate and non-cognate nucleases by Im proteins can vary up to 10 orders of magnitude. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 7.12e-211 | 55 | 538 | 39 | 525 | |
| 1.65e-208 | 56 | 541 | 40 | 522 | |
| 1.30e-207 | 56 | 544 | 63 | 549 | |
| 1.30e-207 | 56 | 544 | 63 | 549 | |
| 1.02e-206 | 56 | 544 | 62 | 548 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.96e-185 | 56 | 538 | 5 | 473 | Penicillium citrinum alpha-1,2-mannosidase complex with glycerol [Penicillium citrinum],2RI8_B Penicillium citrinum alpha-1,2-mannosidase complex with glycerol [Penicillium citrinum],2RI9_A Penicillium citrinum alpha-1,2-mannosidase in complex with a substrate analog [Penicillium citrinum],2RI9_B Penicillium citrinum alpha-1,2-mannosidase in complex with a substrate analog [Penicillium citrinum] |
|
| 1.38e-184 | 56 | 538 | 40 | 508 | Structure of P. citrinum alpha 1,2-mannosidase reveals the basis for differences in specificity of the ER and Golgi Class I enzymes [Penicillium citrinum],1KKT_B Structure of P. citrinum alpha 1,2-mannosidase reveals the basis for differences in specificity of the ER and Golgi Class I enzymes [Penicillium citrinum],1KRE_A Structure Of P. Citrinum Alpha 1,2-Mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum],1KRE_B Structure Of P. Citrinum Alpha 1,2-Mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum],1KRF_A Structure Of P. Citrinum Alpha 1,2-mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum],1KRF_B Structure Of P. Citrinum Alpha 1,2-mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum] |
|
| 7.21e-175 | 59 | 538 | 18 | 492 | Chain A, ALPHA-1,2-MANNOSIDASE [Trichoderma reesei],1HCU_B Chain B, ALPHA-1,2-MANNOSIDASE [Trichoderma reesei],1HCU_C Chain C, ALPHA-1,2-MANNOSIDASE [Trichoderma reesei],1HCU_D Chain D, ALPHA-1,2-MANNOSIDASE [Trichoderma reesei] |
|
| 3.26e-61 | 55 | 538 | 21 | 467 | Structure of mouse Golgi alpha-1,2-mannosidase IA and Man9GlcNAc2-PA complex [Mus musculus],5KKB_B Structure of mouse Golgi alpha-1,2-mannosidase IA and Man9GlcNAc2-PA complex [Mus musculus] |
|
| 4.01e-61 | 55 | 538 | 19 | 465 | Structure of mouse Golgi alpha-1,2-mannosidase IA reveals the molecular basis for substrate specificity among Class I enzymes (family 47 glycosidases) [Mus musculus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.51e-184 | 35 | 538 | 1 | 507 | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=mns1B PE=3 SV=1 |
|
| 7.09e-184 | 56 | 538 | 40 | 508 | Mannosyl-oligosaccharide alpha-1,2-mannosidase OS=Penicillium citrinum OX=5077 GN=MSDC PE=1 SV=2 |
|
| 1.04e-180 | 50 | 538 | 34 | 508 | Mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=mns1B PE=1 SV=1 |
|
| 1.04e-180 | 50 | 538 | 34 | 508 | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=mns1B PE=3 SV=2 |
|
| 9.29e-180 | 47 | 538 | 32 | 510 | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=mns1B PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
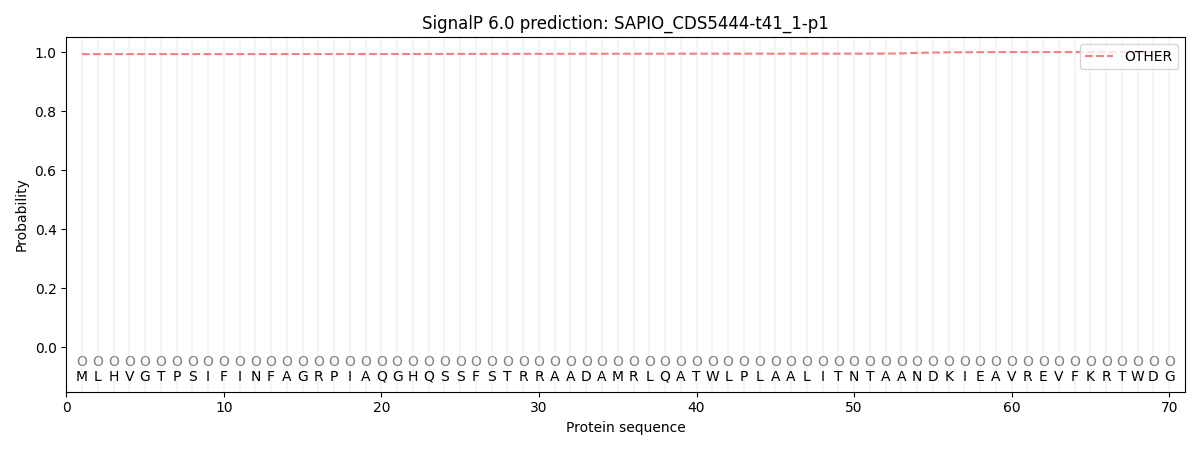
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.993601 | 0.006398 |
