You are browsing environment: FUNGIDB
CAZyme Information: RQM16497.1
You are here: Home > Sequence: RQM16497.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Peronospora effusa | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Peronospora; Peronospora effusa | |||||||||||
| CAZyme ID | RQM16497.1 | |||||||||||
| CAZy Family | GT4 | |||||||||||
| CAZyme Description | 1,3-beta-glucan synthase [Source:UniProtKB/TrEMBL;Acc:A0A3R7WSC9] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.34:28 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT48 | 1114 | 1838 | 8.7e-280 | 0.9553450608930988 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396784 | Glucan_synthase | 0.0 | 1114 | 1879 | 3 | 818 | 1,3-beta-glucan synthase component. This family consists of various 1,3-beta-glucan synthase components including Gls1, Gls2 and Gls3 from yeast. 1,3-beta-glucan synthase EC:2.4.1.34 also known as callose synthase catalyzes the formation of a beta-1,3-glucan polymer that is a major component of the fungal cell wall. The reaction catalyzed is:- UDP-glucose + {(1,3)-beta-D-glucosyl}(N) <=> UDP + {(1,3)-beta-D-glucosyl}(N+1). |
| 405046 | FKS1_dom1 | 1.41e-25 | 235 | 328 | 3 | 106 | 1,3-beta-glucan synthase subunit FKS1, domain-1. The FKS1_dom1 domain is likely to be the 'Class I' region just N-terminal to the first set of transmembrane helices that is involved in 1,3-beta-glucan synthesis itself. This family is found on proteins with family Glucan_synthase, pfam02364. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 2026 | 2045 | 4070 | |
| 0.0 | 2 | 2026 | 18 | 2019 | |
| 0.0 | 2 | 2026 | 17 | 2023 | |
| 2.47e-262 | 123 | 2007 | 82 | 1760 | |
| 1.14e-260 | 132 | 2007 | 68 | 1747 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.79e-253 | 130 | 2003 | 239 | 1917 | Callose synthase 5 OS=Arabidopsis thaliana OX=3702 GN=CALS5 PE=1 SV=1 |
|
| 2.23e-241 | 111 | 2003 | 214 | 1922 | Callose synthase 7 OS=Arabidopsis thaliana OX=3702 GN=CALS7 PE=3 SV=3 |
|
| 5.01e-237 | 130 | 2003 | 262 | 1896 | Callose synthase 10 OS=Arabidopsis thaliana OX=3702 GN=CALS10 PE=2 SV=5 |
|
| 9.67e-237 | 110 | 2001 | 202 | 1936 | Callose synthase 1 OS=Arabidopsis thaliana OX=3702 GN=CALS1 PE=1 SV=2 |
|
| 2.88e-236 | 110 | 2001 | 206 | 1941 | Callose synthase 3 OS=Arabidopsis thaliana OX=3702 GN=CALS3 PE=3 SV=3 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000049 | 0.000000 |
TMHMM Annotations download full data without filtering help
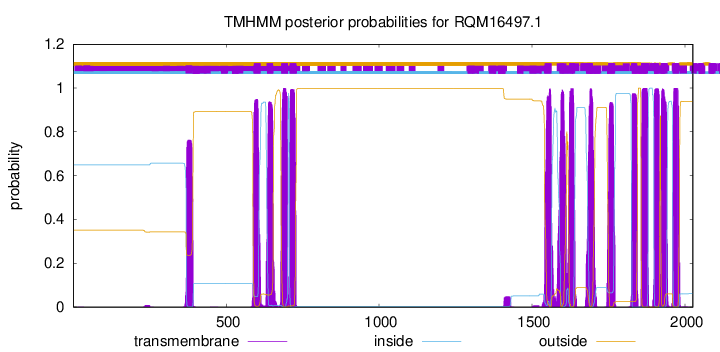
| Start | End |
|---|---|
| 591 | 613 |
| 634 | 653 |
| 680 | 702 |
| 707 | 729 |
| 1542 | 1564 |
| 1590 | 1612 |
| 1622 | 1641 |
| 1677 | 1699 |
| 1750 | 1772 |
| 1827 | 1844 |
| 1857 | 1879 |
| 1899 | 1916 |
| 1921 | 1943 |
| 1960 | 1982 |
