You are browsing environment: FUNGIDB
CAZyme Information: RQM13478.1
You are here: Home > Sequence: RQM13478.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Peronospora effusa | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Peronospora; Peronospora effusa | |||||||||||
| CAZyme ID | RQM13478.1 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 844 | 1183 | 3.9e-147 | 0.997093023255814 |
| GH5 | 265 | 607 | 3.1e-144 | 0.997093023255814 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 225344 | BglC | 2.30e-21 | 841 | 1207 | 39 | 391 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| 225344 | BglC | 1.58e-13 | 262 | 636 | 39 | 383 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| 395098 | Cellulase | 4.74e-10 | 866 | 1183 | 17 | 272 | Cellulase (glycosyl hydrolase family 5). |
| 395098 | Cellulase | 3.07e-07 | 287 | 607 | 17 | 272 | Cellulase (glycosyl hydrolase family 5). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 48 | 1238 | 1 | 1227 | |
| 0.0 | 664 | 1233 | 3 | 575 | |
| 0.0 | 664 | 1233 | 3 | 575 | |
| 0.0 | 664 | 1236 | 3 | 577 | |
| 0.0 | 664 | 1236 | 3 | 577 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.53e-42 | 841 | 1205 | 22 | 361 | Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],3W6M_B Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],3W6M_C Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM2_A Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM2_B Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM2_C Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3] |
|
| 4.78e-42 | 841 | 1205 | 22 | 361 | The Crystal Structure of Cellulase-Inhibitor Complex. [Pyrococcus horikoshii OT3],3VVG_B The Crystal Structure of Cellulase-Inhibitor Complex. [Pyrococcus horikoshii OT3],3VVG_C The Crystal Structure of Cellulase-Inhibitor Complex. [Pyrococcus horikoshii OT3],3W6L_A Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],3W6L_B Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],3W6L_C Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM1_A Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM1_B Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM1_C Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3] |
|
| 2.64e-41 | 841 | 1205 | 55 | 394 | Functional Analysis of Hyperthermophilic Endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3],2ZUN_A Functional Analysis of Hyperthermophilic Endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3],2ZUN_B Functional Analysis of Hyperthermophilic Endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3],2ZUN_C Functional Analysis of Hyperthermophilic Endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3] |
|
| 3.57e-41 | 841 | 1205 | 55 | 394 | Functional analysis of hyperthermophilic endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3],3AXX_B Functional analysis of hyperthermophilic endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3],3AXX_C Functional analysis of hyperthermophilic endocellulase from the Archaeon Pyrococcus horikoshii [Pyrococcus horikoshii OT3] |
|
| 1.18e-40 | 841 | 1205 | 55 | 394 | Crystal analysis of the complex structure, Y299F-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii],3QHO_B Crystal analysis of the complex structure, Y299F-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii],3QHO_C Crystal analysis of the complex structure, Y299F-cellotetraose, of endocellulase from pyrococcus horikoshii [Pyrococcus horikoshii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.35e-28 | 249 | 634 | 39 | 372 | Endoglucanase OS=Paenibacillus polymyxa OX=1406 PE=3 SV=2 |
|
| 2.45e-24 | 828 | 1205 | 46 | 377 | Endoglucanase E1 OS=Acidothermus cellulolyticus (strain ATCC 43068 / DSM 8971 / 11B) OX=351607 GN=Acel_0614 PE=1 SV=1 |
|
| 4.05e-22 | 847 | 1211 | 42 | 360 | Major extracellular endoglucanase OS=Xanthomonas campestris pv. campestris (strain ATCC 33913 / DSM 3586 / NCPPB 528 / LMG 568 / P 25) OX=190485 GN=engXCA PE=1 SV=2 |
|
| 7.92e-19 | 259 | 519 | 638 | 907 | Endoglucanase/exoglucanase B OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=celB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
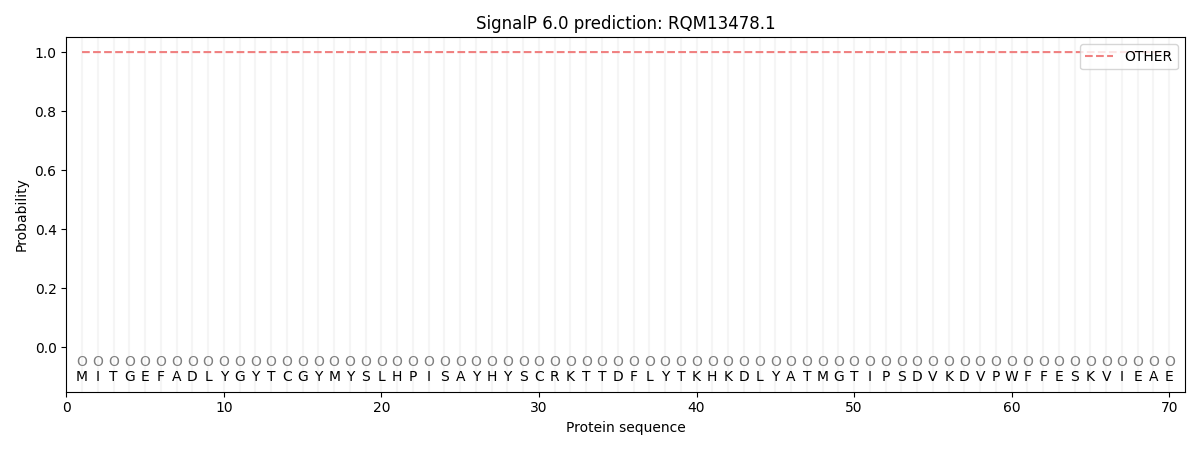
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000057 | 0.000002 |

