You are browsing environment: FUNGIDB
CAZyme Information: RQM12341.1
You are here: Home > Sequence: RQM12341.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Peronospora effusa | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Peronospora; Peronospora effusa | |||||||||||
| CAZyme ID | RQM12341.1 | |||||||||||
| CAZy Family | GH17 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH17 | 45 | 293 | 1.7e-22 | 0.9163987138263665 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 227625 | Scw11 | 1.66e-48 | 17 | 290 | 46 | 304 | Exo-beta-1,3-glucanase, GH17 family [Carbohydrate transport and metabolism]. |
| 366033 | Glyco_hydro_17 | 1.32e-12 | 18 | 294 | 2 | 304 | Glycosyl hydrolases family 17. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.24e-256 | 1 | 363 | 1 | 363 | |
| 1.05e-167 | 1 | 315 | 13 | 327 | |
| 7.77e-165 | 1 | 342 | 13 | 347 | |
| 3.15e-74 | 18 | 294 | 22 | 304 | |
| 2.52e-68 | 2 | 294 | 11 | 303 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.58e-26 | 18 | 290 | 39 | 285 | Crystal structure of glycoside hydrolase family 17 beta-1,3-glucanosyltransferase from Rhizomucor miehei [Rhizomucor miehei CAU432] |
|
| 2.49e-25 | 18 | 290 | 39 | 285 | Active-site mutant of Rhizomucor miehei beta-1,3-glucanosyltransferase in complex with laminaribiose [Rhizomucor miehei CAU432],4WTS_A Active-site mutant of Rhizomucor miehei beta-1,3-glucanosyltransferase in complex with laminaritriose [Rhizomucor miehei CAU432] |
|
| 6.29e-08 | 171 | 294 | 230 | 380 | Crystal structure of an endo-laminarinase from Formosa Hel1_33_131 [Formosa sp. Hel1_33_131],6FCG_B Crystal structure of an endo-laminarinase from Formosa Hel1_33_131 [Formosa sp. Hel1_33_131],6FCG_C Crystal structure of an endo-laminarinase from Formosa Hel1_33_131 [Formosa sp. Hel1_33_131],6FCG_D Crystal structure of an endo-laminarinase from Formosa Hel1_33_131 [Formosa sp. Hel1_33_131],6FCG_E Crystal structure of an endo-laminarinase from Formosa Hel1_33_131 [Formosa sp. Hel1_33_131],6FCG_F Crystal structure of an endo-laminarinase from Formosa Hel1_33_131 [Formosa sp. Hel1_33_131] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.46e-27 | 32 | 302 | 36 | 313 | Glucan 1,3-beta-glucosidase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=BGL2 PE=1 SV=1 |
|
| 9.90e-23 | 33 | 302 | 32 | 308 | Glucan 1,3-beta-glucosidase OS=Candida albicans OX=5476 GN=BGL2 PE=3 SV=1 |
|
| 1.36e-22 | 33 | 302 | 32 | 308 | Glucan 1,3-beta-glucosidase BGL2 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=BGL2 PE=1 SV=2 |
|
| 1.87e-22 | 33 | 302 | 34 | 304 | Glucan 1,3-beta-glucosidase ARB_02797 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_02797 PE=1 SV=1 |
|
| 1.12e-19 | 29 | 295 | 359 | 642 | Probable glucan endo-1,3-beta-glucosidase btgC OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=btgC PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
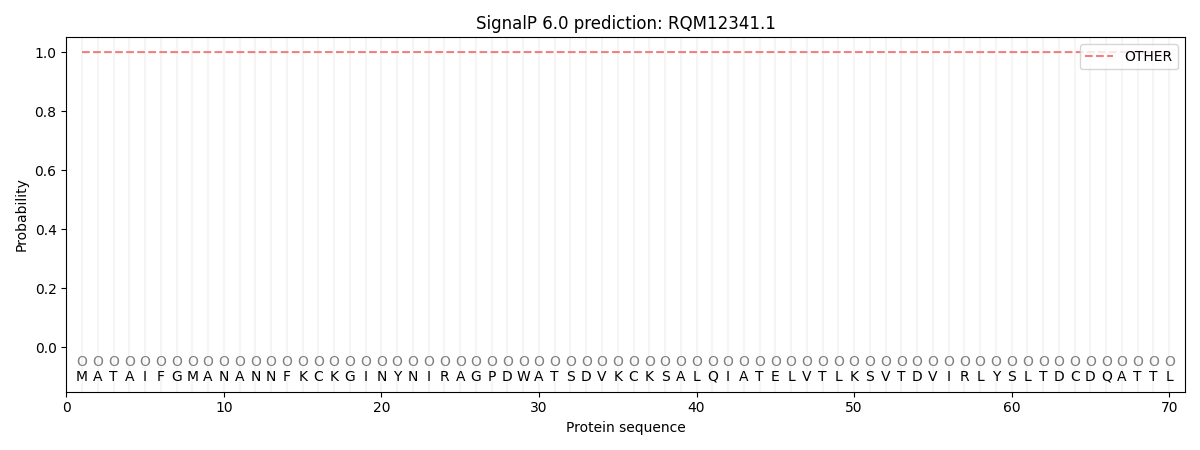
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000033 | 0.000021 |
