You are browsing environment: FUNGIDB
CAZyme Information: RO3G_06021-t26_1-p1
You are here: Home > Sequence: RO3G_06021-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Rhizopus delemar | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Mucoromycota; Mucoromycetes; ; Rhizopodaceae; Rhizopus; Rhizopus delemar | |||||||||||
| CAZyme ID | RO3G_06021-t26_1-p1 | |||||||||||
| CAZy Family | GH27 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 50 | 376 | 2.7e-70 | 0.9661538461538461 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395231 | Glyco_hydro_28 | 2.43e-48 | 52 | 370 | 1 | 315 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| 215426 | PLN02793 | 3.05e-46 | 35 | 377 | 60 | 412 | Probable polygalacturonase |
| 177865 | PLN02218 | 7.57e-46 | 35 | 384 | 75 | 431 | polygalacturonase ADPG |
| 215540 | PLN03010 | 1.03e-42 | 35 | 383 | 54 | 402 | polygalacturonase |
| 178580 | PLN03003 | 7.89e-40 | 39 | 380 | 37 | 381 | Probable polygalacturonase At3g15720 |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.01e-149 | 1 | 384 | 1 | 382 | |
| 4.01e-149 | 1 | 384 | 1 | 382 | |
| 5.56e-108 | 12 | 383 | 8 | 379 | |
| 4.15e-88 | 11 | 383 | 7 | 395 | |
| 9.18e-61 | 40 | 383 | 62 | 432 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.62e-33 | 37 | 373 | 27 | 373 | Crystal structure of endo-xylogalacturonan hydrolase from Aspergillus tubingensis [Aspergillus tubingensis] |
|
| 2.20e-27 | 104 | 374 | 63 | 331 | Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_B Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_C Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_D Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_E Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_F Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_G Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini] |
|
| 5.75e-27 | 104 | 374 | 67 | 333 | Polygalacturonase From Aspergillus Aculeatus [Aspergillus aculeatus],1IB4_A Crystal Structure of Polygalacturonase from Aspergillus Aculeatus at Ph4.5 [Aspergillus aculeatus],1IB4_B Crystal Structure of Polygalacturonase from Aspergillus Aculeatus at Ph4.5 [Aspergillus aculeatus] |
|
| 3.49e-25 | 107 | 370 | 68 | 330 | Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_B Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_C Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_D Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_E Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_F Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger] |
|
| 6.81e-24 | 104 | 368 | 71 | 334 | Chain A, Endo-polygalacturonase [Evansstolkia leycettana] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.06e-284 | 1 | 385 | 1 | 385 | Exopolygalacturonase rpg16 OS=Rhizopus delemar (strain RA 99-880 / ATCC MYA-4621 / FGSC 9543 / NRRL 43880) OX=246409 GN=rpg16 PE=1 SV=1 |
|
| 6.58e-215 | 1 | 385 | 1 | 385 | Exopolygalacturonase rpg15 OS=Rhizopus delemar (strain RA 99-880 / ATCC MYA-4621 / FGSC 9543 / NRRL 43880) OX=246409 GN=rpg15 PE=1 SV=1 |
|
| 1.38e-142 | 1 | 385 | 1 | 363 | Exopolygalacturonase rpg13 OS=Rhizopus delemar (strain RA 99-880 / ATCC MYA-4621 / FGSC 9543 / NRRL 43880) OX=246409 GN=rpg13 PE=1 SV=1 |
|
| 1.01e-139 | 1 | 384 | 1 | 372 | Exopolygalacturonase rpg14 OS=Rhizopus delemar (strain RA 99-880 / ATCC MYA-4621 / FGSC 9543 / NRRL 43880) OX=246409 GN=rpg14 PE=1 SV=1 |
|
| 1.05e-58 | 25 | 385 | 44 | 433 | Probable exopolygalacturonase X OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=pgaX PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
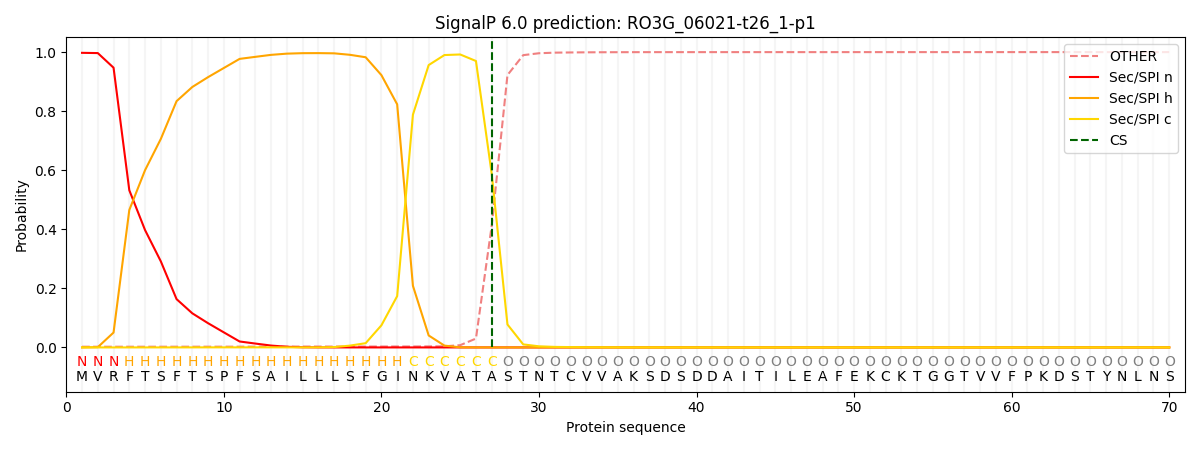
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.003580 | 0.996360 | CS pos: 27-28. Pr: 0.5877 |
