You are browsing environment: FUNGIDB
CAZyme Information: RL4_JR_09875-RA-p1
You are here: Home > Sequence: RL4_JR_09875-RA-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Raffaelea lauricola | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Ophiostomataceae; Raffaelea; Raffaelea lauricola | |||||||||||
| CAZyme ID | RL4_JR_09875-RA-p1 | |||||||||||
| CAZy Family | GT4 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.-:5 | 3.2.1.21:4 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 103 | 321 | 1.3e-58 | 0.9814814814814815 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 185053 | PRK15098 | 4.57e-58 | 127 | 805 | 118 | 758 | beta-glucosidase BglX. |
| 396478 | Glyco_hydro_3_C | 2.64e-54 | 424 | 672 | 1 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| 224389 | BglX | 1.34e-45 | 121 | 455 | 75 | 374 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| 178629 | PLN03080 | 3.58e-37 | 72 | 798 | 52 | 769 | Probable beta-xylosidase; Provisional |
| 395747 | Glyco_hydro_3 | 5.23e-34 | 126 | 364 | 85 | 315 | Glycosyl hydrolase family 3 N terminal domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 3 | 815 | 2 | 815 | |
| 0.0 | 3 | 815 | 2 | 815 | |
| 0.0 | 3 | 815 | 2 | 815 | |
| 0.0 | 3 | 815 | 2 | 815 | |
| 0.0 | 3 | 815 | 2 | 815 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.33e-197 | 50 | 810 | 24 | 851 | Chain A, Beta-glucosidase [Rasamsonia emersonii],5JU6_B Chain B, Beta-glucosidase [Rasamsonia emersonii],5JU6_C Chain C, Beta-glucosidase [Rasamsonia emersonii],5JU6_D Chain D, Beta-glucosidase [Rasamsonia emersonii] |
|
| 4.83e-194 | 62 | 810 | 2 | 710 | Crystasl Structure of Beta-glucosidase D2-BGL from Chaetomella Raphigera [Chaetomella raphigera] |
|
| 1.84e-191 | 38 | 804 | 10 | 850 | Structural studies of a Glycoside Hydrolase Family 3 beta-glucosidase from the Model Fungus Neurospora crassa [Neurospora crassa OR74A],5NBS_B Structural studies of a Glycoside Hydrolase Family 3 beta-glucosidase from the Model Fungus Neurospora crassa [Neurospora crassa OR74A] |
|
| 1.16e-189 | 66 | 811 | 9 | 711 | Chain A, Beta-d-glucoside Glucohydrolase [Trichoderma reesei],3ZZ1_A Chain A, BETA-D-GLUCOSIDE GLUCOHYDROLASE [Trichoderma reesei] |
|
| 1.19e-189 | 66 | 811 | 10 | 712 | Chain A, Beta-D-glucoside glucohydrolase [Trichoderma reesei],4I8D_B Chain B, Beta-D-glucoside glucohydrolase [Trichoderma reesei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 3 | 811 | 2 | 811 | Probable beta-glucosidase G OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=bglG PE=3 SV=1 |
|
| 0.0 | 3 | 805 | 2 | 805 | Probable beta-glucosidase G OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=bglG PE=3 SV=1 |
|
| 0.0 | 3 | 815 | 2 | 815 | Probable beta-glucosidase G OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=bglG PE=3 SV=1 |
|
| 0.0 | 3 | 815 | 2 | 815 | Probable beta-glucosidase G OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=bglG PE=3 SV=1 |
|
| 0.0 | 26 | 811 | 26 | 815 | Probable beta-glucosidase G OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=bglG PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
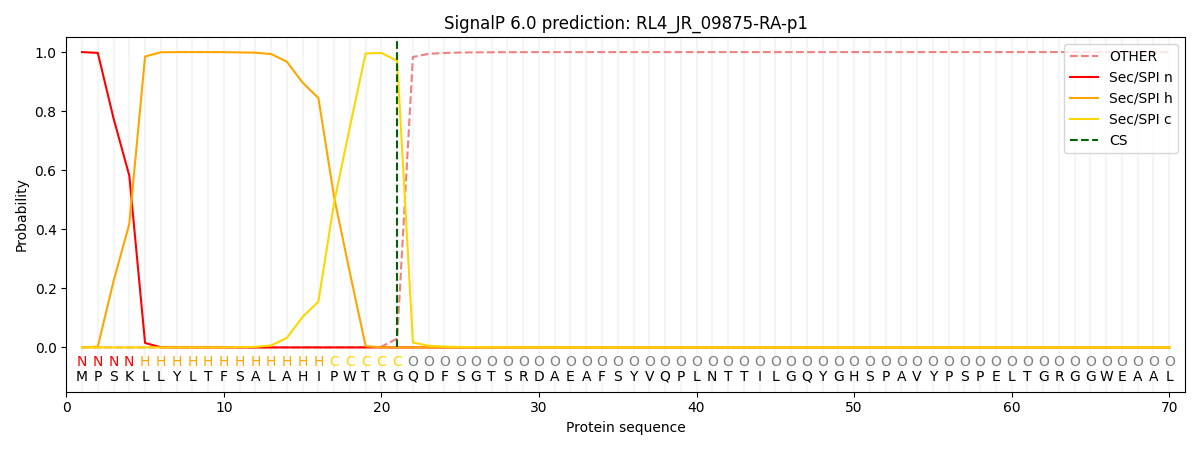
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000263 | 0.999705 | CS pos: 21-22. Pr: 0.9703 |
