You are browsing environment: FUNGIDB
CAZyme Information: RL4_JR_01530-RA-p1
You are here: Home > Sequence: RL4_JR_01530-RA-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Raffaelea lauricola | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Ophiostomataceae; Raffaelea; Raffaelea lauricola | |||||||||||
| CAZyme ID | RL4_JR_01530-RA-p1 | |||||||||||
| CAZy Family | AA4 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT22 | 4 | 380 | 1.5e-70 | 0.9460154241645244 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 281842 | Glyco_transf_22 | 2.14e-45 | 3 | 379 | 17 | 410 | Alg9-like mannosyltransferase family. Members of this family are mannosyltransferase enzymes. At least some members are localized in endoplasmic reticulum and involved in GPI anchor biosynthesis. |
| 215437 | PLN02816 | 0.003 | 232 | 362 | 266 | 426 | mannosyltransferase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.54e-293 | 1 | 503 | 18 | 520 | |
| 1.74e-291 | 1 | 500 | 18 | 517 | |
| 1.04e-288 | 1 | 498 | 18 | 515 | |
| 3.74e-287 | 1 | 500 | 18 | 517 | |
| 5.12e-287 | 1 | 498 | 18 | 515 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.81e-278 | 1 | 500 | 18 | 517 | GPI mannosyltransferase 4 OS=Gibberella zeae (strain ATCC MYA-4620 / CBS 123657 / FGSC 9075 / NRRL 31084 / PH-1) OX=229533 GN=SMP3 PE=3 SV=2 |
|
| 1.57e-169 | 1 | 497 | 18 | 540 | GPI mannosyltransferase 4 OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=smp3 PE=3 SV=1 |
|
| 1.30e-161 | 1 | 492 | 18 | 539 | GPI mannosyltransferase 4 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=smp3 PE=3 SV=2 |
|
| 1.11e-154 | 1 | 492 | 18 | 534 | GPI mannosyltransferase 4 OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=smp3 PE=3 SV=2 |
|
| 6.01e-91 | 1 | 470 | 21 | 495 | GPI mannosyltransferase 4 OS=Yarrowia lipolytica (strain CLIB 122 / E 150) OX=284591 GN=SMP3 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999742 | 0.000255 |
TMHMM Annotations download full data without filtering help
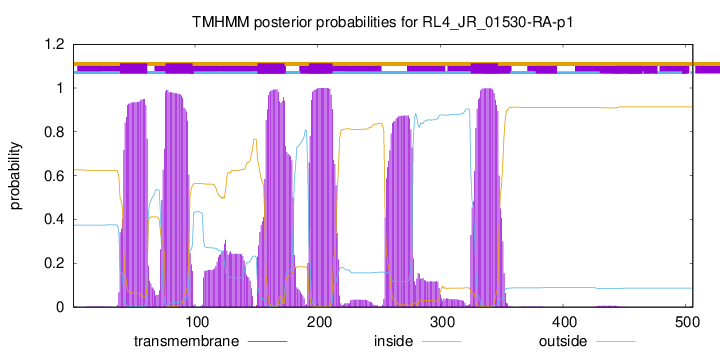
| Start | End |
|---|---|
| 39 | 61 |
| 76 | 98 |
| 151 | 173 |
| 193 | 212 |
| 325 | 347 |
