You are browsing environment: FUNGIDB
CAZyme Information: RAK81459.1
You are here: Home > Sequence: RAK81459.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus fijiensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus fijiensis | |||||||||||
| CAZyme ID | RAK81459.1 | |||||||||||
| CAZy Family | GT24 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH16 | 99 | 270 | 4.7e-72 | 0.9770114942528736 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 185692 | GH16_fungal_CRH1_transglycosylase | 4.08e-90 | 86 | 295 | 3 | 203 | glycosylphosphatidylinositol-glucanosyltransferase. Group of fungal GH16 members related to Saccharomyces cerevisiae Crh1p. Chr1p and Crh2p are transglycosylases that are required for the linkage of chitin to beta(1-3)glucose branches of beta(1-6)glucan, an important step in the assembly of new cell wall. Both have been shown to be glycosylphosphatidylinositol (GPI)-anchored. A third homologous protein, Crr1p, functions in the formation of the spore wall. They belongs to the family 16 of glycosyl hydrolases that includes lichenase, xyloglucan endotransglycosylase (XET), beta-agarase, kappa-carrageenase, endo-beta-1,3-glucanase, endo-beta-1,3-1,4-glucanase, and endo-beta-galactosidase, all of which have a conserved jelly roll fold with a deep active site channel harboring the catalytic residues. |
| 395585 | Glyco_hydro_16 | 1.38e-48 | 97 | 264 | 1 | 166 | Glycosyl hydrolases family 16. |
| 185683 | Glyco_hydrolase_16 | 1.59e-18 | 111 | 271 | 49 | 206 | glycosyl hydrolase family 16. The O-Glycosyl hydrolases are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A glycosyl hydrolase classification system based on sequence similarity has led to the definition of more than 95 different families inlcuding glycosyl hydrolase family 16. Family 16 includes lichenase, xyloglucan endotransglycosylase (XET), beta-agarase, kappa-carrageenase, endo-beta-1,3-glucanase, endo-beta-1,3-1,4-glucanase, and endo-beta-galactosidase, all of which have a conserved jelly roll fold with a deep active site channel harboring the catalytic residues. |
| 185684 | GH16_lichenase | 1.95e-18 | 102 | 266 | 35 | 193 | lichenase, member of glycosyl hydrolase family 16. Lichenase, also known as 1,3-1,4-beta-glucanase, is a member of glycosyl hydrolase family 16, that specifically cleaves 1,4-beta-D-glucosidic bonds in mixed-linked beta glucans that also contain 1,3-beta-D-glucosidic linkages. Natural substrates of beta-glucanase are beta-glucans from grain endosperm cell walls or lichenan from the Islandic moss, Cetraria islandica. This protein is found not only in bacteria but also in anaerobic fungi. This domain includes two seven-stranded antiparallel beta-sheets that are adjacent to one another forming a compact, jellyroll beta-sandwich structure. |
| 225182 | BglS | 6.79e-18 | 28 | 279 | 18 | 265 | Beta-glucanase, GH16 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.52e-190 | 1 | 353 | 1 | 355 | |
| 5.63e-189 | 1 | 355 | 1 | 357 | |
| 5.63e-189 | 1 | 355 | 1 | 357 | |
| 4.33e-137 | 1 | 337 | 2 | 339 | |
| 4.68e-134 | 1 | 354 | 2 | 361 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.16e-31 | 50 | 315 | 4 | 237 | Apo Crh5 transglycosylase [Aspergillus fumigatus Af293],6IBU_B Apo Crh5 transglycosylase [Aspergillus fumigatus Af293],6IBW_A Crh5 transglycosylase in complex with NAG [Aspergillus fumigatus Af293],6IBW_B Crh5 transglycosylase in complex with NAG [Aspergillus fumigatus Af293] |
|
| 1.22e-15 | 64 | 264 | 8 | 192 | Bacillus Licheniformis Beta-Glucanase [Bacillus licheniformis] |
|
| 6.12e-15 | 129 | 264 | 59 | 179 | Crystal structure of the engineered 1,3-1,4-beta-glucanase protein from Bacillus licheniformis [Bacillus licheniformis],3D6E_B Crystal structure of the engineered 1,3-1,4-beta-glucanase protein from Bacillus licheniformis [Bacillus licheniformis] |
|
| 4.54e-13 | 129 | 264 | 96 | 216 | Crystal Structure of the endo-beta-1,3-1,4 glucanase from Bacillus subtilis (strain 168) [Bacillus subtilis] |
|
| 4.42e-12 | 129 | 275 | 14 | 142 | NATIVE-LIKE IN VIVO FOLDING OF A CIRCULARLY PERMUTED JELLYROLL PROTEIN SHOWN BY CRYSTAL STRUCTURE ANALYSIS [Paenibacillus macerans] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.17e-139 | 1 | 352 | 2 | 359 | Probable glycosidase crf2 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=crf2 PE=1 SV=1 |
|
| 2.26e-97 | 5 | 336 | 4 | 352 | Extracellular glycosidase UTR2 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=UTR2 PE=1 SV=1 |
|
| 1.14e-90 | 18 | 348 | 23 | 356 | Probable glycosidase CRH2 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=UTR2 PE=1 SV=3 |
|
| 8.02e-54 | 22 | 295 | 31 | 363 | Probable glycosidase CRR1 OS=Ashbya gossypii (strain ATCC 10895 / CBS 109.51 / FGSC 9923 / NRRL Y-1056) OX=284811 GN=CRR1 PE=3 SV=1 |
|
| 2.30e-53 | 19 | 294 | 28 | 354 | Probable glycosidase CRR1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=CRR1 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
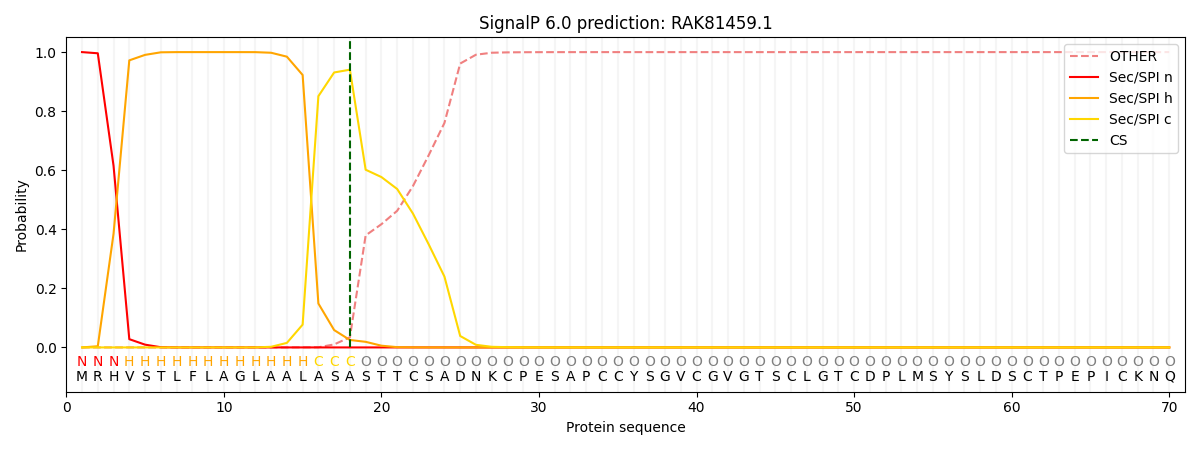
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000296 | 0.999668 | CS pos: 18-19. Pr: 0.9404 |
