You are browsing environment: FUNGIDB
CAZyme Information: RAK71145.1
You are here: Home > Sequence: RAK71145.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus fijiensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus fijiensis | |||||||||||
| CAZyme ID | RAK71145.1 | |||||||||||
| CAZy Family | AA1 | |||||||||||
| CAZyme Description | endo-1,4-beta-xylanase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.8:67 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH11 | 50 | 229 | 1.5e-72 | 0.9943502824858758 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395367 | Glyco_hydro_11 | 1.33e-99 | 50 | 227 | 1 | 175 | Glycosyl hydrolases family 11. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.54e-158 | 1 | 231 | 1 | 231 | |
| 3.69e-157 | 1 | 231 | 1 | 231 | |
| 2.87e-153 | 1 | 231 | 1 | 233 | |
| 2.87e-153 | 1 | 231 | 1 | 233 | |
| 3.94e-103 | 4 | 230 | 2 | 224 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.97e-76 | 53 | 231 | 15 | 190 | Crystal structure of family 11 xylanase in complex with inhibitor (XIP-I) [Talaromyces funiculosus] |
|
| 2.23e-75 | 53 | 231 | 32 | 207 | Xylanase 11C from Talaromyces cellulolyticus (formerly known as Acremonium cellulolyticus) [Talaromyces funiculosus],3WP3_B Xylanase 11C from Talaromyces cellulolyticus (formerly known as Acremonium cellulolyticus) [Talaromyces funiculosus] |
|
| 1.48e-73 | 53 | 231 | 39 | 215 | Crystal structure of environmental isolated GH11 in complex with xylobiose and feruloyl-arabino-xylotriose [Escherichia coli] |
|
| 1.53e-73 | 53 | 231 | 39 | 215 | Environmentally isolated GH11 xylanase [Escherichia coli] |
|
| 1.70e-73 | 53 | 231 | 15 | 190 | Chain A, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_B Chain B, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_C Chain C, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_D Chain D, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_E Chain E, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_F Chain F, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_G Chain G, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_H Chain H, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_I Chain I, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_J Chain J, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_K Chain K, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_L Chain L, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.99e-104 | 4 | 230 | 2 | 224 | Endo-1,4-beta-xylanase B OS=Aspergillus kawachii (strain NBRC 4308) OX=1033177 GN=xlnB PE=3 SV=2 |
|
| 4.68e-102 | 4 | 230 | 2 | 224 | Endo-1,4-beta-xylanase B OS=Aspergillus niger OX=5061 GN=xlnB PE=1 SV=1 |
|
| 4.68e-102 | 4 | 230 | 2 | 224 | Probable endo-1,4-beta-xylanase B OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=xlnB PE=3 SV=1 |
|
| 1.39e-100 | 1 | 230 | 1 | 231 | Probable endo-1,4-beta-xylanase A OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=xlnA PE=3 SV=2 |
|
| 1.39e-100 | 1 | 230 | 1 | 231 | Endo-1,4-beta-xylanase A OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=xlnA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
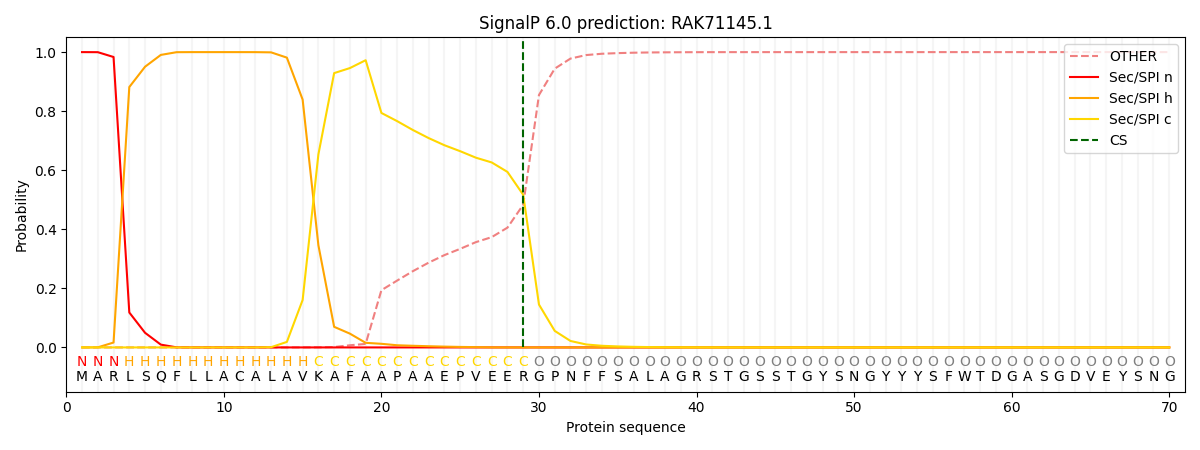
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000211 | 0.999749 | CS pos: 29-30. Pr: 0.5168 |
