You are browsing environment: FUNGIDB
CAZyme Information: QYA_121587T0-p1
You are here: Home > Sequence: QYA_121587T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Mucor lusitanicus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Mucoromycota; Mucoromycetes; ; Mucoraceae; Mucor; Mucor lusitanicus | |||||||||||
| CAZyme ID | QYA_121587T0-p1 | |||||||||||
| CAZy Family | CE9 | |||||||||||
| CAZyme Description | Chitinase [Source:UniProtKB/TrEMBL;Acc:A0A168H2G1] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.14:12 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH18 | 1 | 236 | 7.1e-17 | 0.6418918918918919 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 119356 | GH18_hevamine_XipI_class_III | 6.99e-80 | 1 | 272 | 3 | 263 | This conserved domain family includes xylanase inhibitor Xip-I, and the class III plant chitinases such as hevamine, concanavalin B, and PPL2, all of which have a glycosyl hydrolase family 18 (GH18) domain. Hevamine is a class III endochitinase that hydrolyzes the linear polysaccharide chains of chitin and peptidoglycan and is important for defense against pathogenic bacteria and fungi. PPL2 (Parkia platycephala lectin 2) is a class III chitinase from Parkia platycephala seeds that hydrolyzes beta(1-4) glycosidic bonds linking 2-acetoamido-2-deoxy-beta-D-glucopyranose units in chitin. |
| 395573 | Glyco_hydro_18 | 1.47e-11 | 1 | 236 | 2 | 220 | Glycosyl hydrolases family 18. |
| 119349 | GH18_chitinase-like | 5.40e-08 | 1 | 197 | 1 | 177 | The GH18 (glycosyl hydrolase, family 18) type II chitinases hydrolyze chitin, an abundant polymer of beta-1,4-linked N-acetylglucosamine (GlcNAc) which is a major component of the cell wall of fungi and the exoskeleton of arthropods. Chitinases have been identified in viruses, bacteria, fungi, protozoan parasites, insects, and plants. The structure of the GH18 domain is an eight-stranded beta/alpha barrel with a pronounced active-site cleft at the C-terminal end of the beta-barrel. The GH18 family includes chitotriosidase, chitobiase, hevamine, zymocin-alpha, narbonin, SI-CLP (stabilin-1 interacting chitinase-like protein), IDGF (imaginal disc growth factor), CFLE (cortical fragment-lytic enzyme) spore hydrolase, the type III and type V plant chitinases, the endo-beta-N-acetylglucosaminidases, and the chitolectins. The GH85 (glycosyl hydrolase, family 85) ENGases (endo-beta-N-acetylglucosaminidases) are closely related to the GH18 chitinases and are included in this alignment model. |
| 119363 | GH18_CTS3_chitinase | 5.90e-07 | 68 | 272 | 69 | 251 | GH18 domain of CTS3 (chitinase 3), an uncharacterized protein from the human fungal pathogen Coccidioides posadasii. CTS3 has a chitinase-like glycosyl hydrolase family 18 (GH18) domain; and has homologs in bacteria as well as fungi. |
| 119350 | GH18_chitinase_D-like | 8.63e-06 | 63 | 200 | 65 | 197 | GH18 domain of Chitinase D (ChiD). ChiD, a chitinase found in Bacillus circulans, hydrolyzes the 1,4-beta-linkages of N-acetylglucosamine in chitin and chitodextrins. The domain architecture of ChiD includes a catalytic glycosyl hydrolase family 18 (GH18) domain, a chitin-binding domain, and a fibronectin type III domain. The chitin-binding and fibronectin type III domains are located either N-terminal or C-terminal to the catalytic domain. This family includes exochitinase Chi36 from Bacillus cereus. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.07e-50 | 1 | 272 | 30 | 287 | |
| 5.22e-50 | 1 | 272 | 27 | 285 | |
| 5.22e-50 | 1 | 272 | 27 | 285 | |
| 5.22e-50 | 1 | 272 | 27 | 285 | |
| 5.22e-50 | 1 | 272 | 27 | 285 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.66e-45 | 1 | 272 | 8 | 264 | ScCTS1_apo crystal structure [Saccharomyces cerevisiae],2UY3_A ScCTS1_8-chlorotheophylline crystal structure [Saccharomyces cerevisiae],2UY4_A ScCTS1_acetazolamide crystal structure [Saccharomyces cerevisiae],2UY5_A ScCTS1_kinetin crystal structure [Saccharomyces cerevisiae],4TXE_A ScCTS1 in complex with compound 5 [Saccharomyces cerevisiae] |
|
| 7.66e-45 | 1 | 272 | 3 | 257 | Crystal structure of class III chitinase from pomegranate provides the insight into its metal storage capacity [Punica granatum],4TOQ_B Crystal structure of class III chitinase from pomegranate provides the insight into its metal storage capacity [Punica granatum],4TOQ_C Crystal structure of class III chitinase from pomegranate provides the insight into its metal storage capacity [Punica granatum],4TOQ_D Crystal structure of class III chitinase from pomegranate provides the insight into its metal storage capacity [Punica granatum] |
|
| 4.34e-40 | 1 | 272 | 3 | 255 | CRYSTAL STRUCTURES OF HEVAMINE, A PLANT DEFENCE PROTEIN WITH CHITINASE AND LYSOZYME ACTIVITY, AND ITS COMPLEX WITH AN INHIBITOR [Hevea brasiliensis],1LLO_A Chain A, HEVAMINE [Hevea brasiliensis],2HVM_A Hevamine A At 1.8 Angstrom Resolution [Hevea brasiliensis] |
|
| 5.80e-40 | 1 | 272 | 3 | 253 | cDNA cloning and 1.75A crystal structure determination of PPL2, a novel chimerolectin from Parkia platycephala seeds exhibiting endochitinolytic activity [Parkia platycephala] |
|
| 2.59e-38 | 1 | 272 | 3 | 255 | Chain A, Hevamine A [Hevea brasiliensis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.98e-46 | 1 | 272 | 31 | 292 | Chitinase 1 OS=Rhizopus oligosporus OX=4847 GN=CHI1 PE=1 SV=1 |
|
| 2.04e-46 | 1 | 272 | 31 | 292 | Chitinase 2 OS=Rhizopus oligosporus OX=4847 GN=CHI2 PE=1 SV=1 |
|
| 2.92e-46 | 1 | 272 | 25 | 287 | Chitinase 3 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=CHT3 PE=1 SV=2 |
|
| 3.04e-45 | 1 | 272 | 28 | 277 | Acidic endochitinase SE2 OS=Beta vulgaris OX=161934 GN=SE2 PE=1 SV=1 |
|
| 3.11e-44 | 1 | 272 | 20 | 270 | Chitinase 1 OS=Candida albicans OX=5476 GN=CHT1 PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
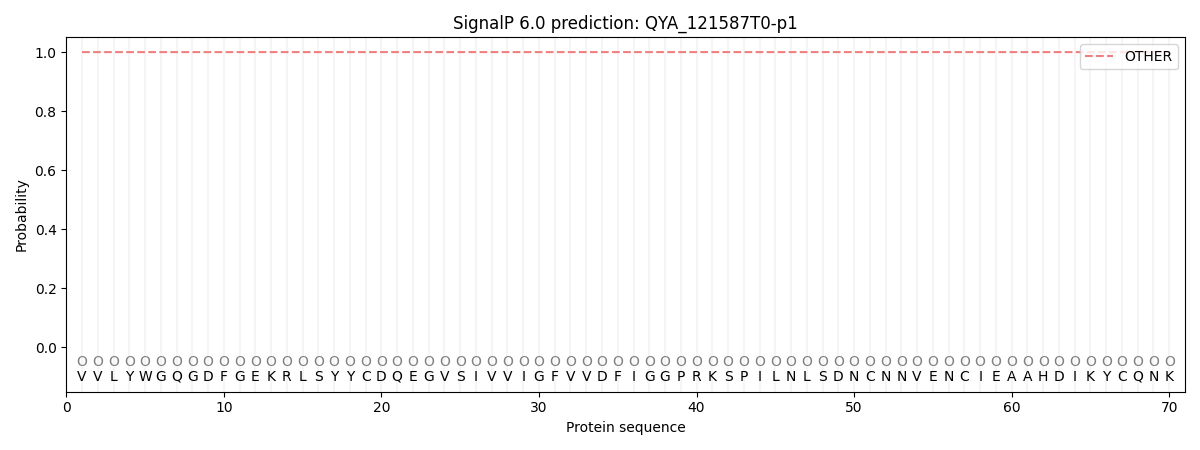
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000021 | 0.000006 |
