You are browsing environment: FUNGIDB
CAZyme Information: QSL66738.1
You are here: Home > Sequence: QSL66738.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Pneumocystis wakefieldiae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Pneumocystidomycetes; ; Pneumocystidaceae; Pneumocystis; Pneumocystis wakefieldiae | |||||||||||
| CAZyme ID | QSL66738.1 | |||||||||||
| CAZy Family | GT58 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.113:99 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 264 | 713 | 1.5e-159 | 0.9932735426008968 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 0.0 | 263 | 713 | 1 | 452 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 7.83e-174 | 256 | 716 | 72 | 520 | glycoside hydrolase family 47 protein; Provisional |
| 239720 | proteasome_alpha_type_3 | 6.43e-125 | 5 | 216 | 1 | 212 | proteasome_alpha_type_3. The 20S proteasome, multisubunit proteolytic complex, is the central enzyme of nonlysosomal protein degradation in both the cytosol and nucleus. It is composed of 28 subunits arranged as four homoheptameric rings that stack on top of one another forming an elongated alpha-beta-beta-alpha cylinder with a central cavity. The proteasome alpha and beta subunits are members of the N-terminal nucleophile (Ntn)-hydrolase superfamily. Their N-terminal threonine residues are exposed as a nucleophile in peptide bond hydrolysis. Mammals have 7 alpha and 7 beta proteasome subunits while archaea have one of each. |
| 238892 | proteasome_alpha | 8.57e-101 | 8 | 216 | 1 | 209 | proteasome alpha subunit. The 20S proteasome, multisubunit proteolytic complex, is the central enzyme of nonlysosomal protein degradation in both the cytosol and nucleus. It is composed of 28 subunits arranged as four homoheptameric rings that stack on top of one another forming an elongated alpha-beta-beta-alpha cylinder with a central cavity. The proteasome alpha and beta subunits are members of the N-terminal nucleophile (Ntn)-hydrolase superfamily. Their N-terminal threonine residues are exposed as a nucleophile in peptide bond hydrolysis. Mammals have 7 different alpha and 10 different beta proteasome subunit genes while archaea have one of each. |
| 235192 | PRK03996 | 5.20e-63 | 6 | 234 | 8 | 238 | archaeal proteasome endopeptidase complex subunit alpha. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 727 | 1 | 727 | |
| 3.94e-184 | 235 | 727 | 101 | 596 | |
| 5.99e-180 | 251 | 726 | 108 | 588 | |
| 5.99e-180 | 251 | 726 | 108 | 588 | |
| 5.99e-180 | 251 | 726 | 108 | 588 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.01e-130 | 257 | 727 | 6 | 510 | Crystal Structure Of Class I Alpha-1,2-mannosidase From Saccharomyces Cerevisiae At 1.54 Angstrom Resolution [Saccharomyces cerevisiae] |
|
| 5.82e-130 | 257 | 727 | 40 | 544 | Crystal structure of the yeast alpha-1,2-mannosidase with bound 1-deoxymannojirimycin at 1.59 A resolution [Saccharomyces cerevisiae] |
|
| 4.01e-125 | 256 | 713 | 5 | 450 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase and Man9GlcNAc2-PA complex [Homo sapiens] |
|
| 4.87e-125 | 256 | 713 | 10 | 455 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase [Homo sapiens] |
|
| 5.19e-125 | 256 | 713 | 10 | 455 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With 1-Deoxymannojirimycin [Homo sapiens],1FO3_A Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With Kifunensine [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.89e-163 | 247 | 727 | 29 | 520 | Mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Candida albicans OX=5476 GN=MNS1 PE=3 SV=2 |
|
| 3.05e-138 | 256 | 716 | 46 | 513 | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPAC2E1P5.01c PE=1 SV=2 |
|
| 4.85e-131 | 250 | 713 | 122 | 619 | Mannosyl-oligosaccharide 1,2-alpha-mannosidase MNS3 OS=Arabidopsis thaliana OX=3702 GN=MNS3 PE=1 SV=1 |
|
| 3.75e-128 | 257 | 727 | 39 | 548 | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=MNS1 PE=1 SV=1 |
|
| 1.02e-121 | 213 | 713 | 164 | 652 | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Rattus norvegicus OX=10116 GN=Man1b1 PE=2 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
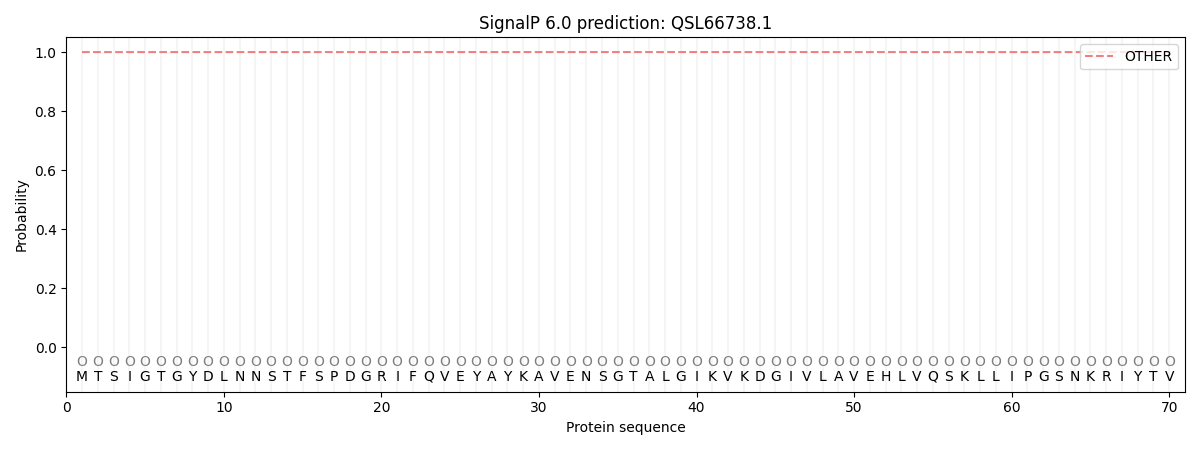
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000079 | 0.000002 |
